Analysis of human C1q by combined bottom-up and top-down mass spectrometry: detailed mapping of post-translational modifications and insights into the C1r/C1s binding sites
- PMID: 20008834
- PMCID: PMC2860232
- DOI: 10.1074/mcp.M900350-MCP200
Analysis of human C1q by combined bottom-up and top-down mass spectrometry: detailed mapping of post-translational modifications and insights into the C1r/C1s binding sites
Abstract
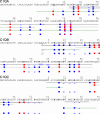
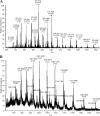
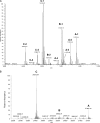
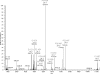
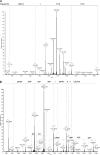
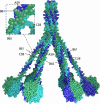
C1q is a subunit of the C1 complex, a key player in innate immunity that triggers activation of the classical complement pathway. Featuring a unique structural organization and comprising a collagen-like domain with a high level of post-translational modifications, C1q represents a challenging protein assembly for structural biology. We report for the first time a comprehensive proteomics study of C1q combining bottom-up and top-down analyses. C1q was submitted to proteolytic digestion by a combination of collagenase and trypsin for bottom-up analyses. In addition to classical LC-MS/MS analyses, which provided reliable identification of hydroxylated proline and lysine residues, sugar loss-triggered MS(3) scans were acquired on an LTQ-Orbitrap (Linear Quadrupole Ion Trap-Orbitrap) instrument to strengthen the localization of glucosyl-galactosyl disaccharide moieties on hydroxylysine residues. Top-down analyses performed on the same instrument allowed high accuracy and high resolution mass measurements of the intact full-length C1q polypeptide chains and the iterative fragmentation of the proteins in the MS(n) mode. This study illustrates the usefulness of combining the two complementary analytical approaches to obtain a detailed characterization of the post-translational modification pattern of the collagen-like domain of C1q and highlights the structural heterogeneity of individual molecules. Most importantly, three lysine residues of the collagen-like domain, namely Lys(59) (A chain), Lys(61) (B chain), and Lys(58) (C chain), were unambiguously shown to be completely unmodified. These lysine residues are located about halfway along the collagen-like fibers. They are thus fully available and in an appropriate position to interact with the C1r and C1s protease partners of C1q and are therefore likely to play an essential role in C1 assembly.
Figures
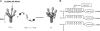



Similar articles
-
Mapping surface accessibility of the C1r/C1s tetramer by chemical modification and mass spectrometry provides new insights into assembly of the human C1 complex.J Biol Chem. 2010 Oct 15;285(42):32251-63. doi: 10.1074/jbc.M110.149112. Epub 2010 Jun 30. J Biol Chem. 2010. PMID: 20592021 Free PMC article.
-
Expression of recombinant human complement C1q allows identification of the C1r/C1s-binding sites.Proc Natl Acad Sci U S A. 2013 May 21;110(21):8650-5. doi: 10.1073/pnas.1304894110. Epub 2013 May 6. Proc Natl Acad Sci U S A. 2013. PMID: 23650384 Free PMC article.
-
Chemical characterization and location of ionic interactions involved in the assembly of the C1 complex of human complement.J Protein Chem. 1993 Dec;12(6):771-81. doi: 10.1007/BF01024936. J Protein Chem. 1993. PMID: 8136028
-
The structure and function of the first component of complement: genetic engineering approach (a review).Acta Microbiol Immunol Hung. 1994;41(4):361-80. Acta Microbiol Immunol Hung. 1994. PMID: 7866721 Review.
-
C1 subcomponent complexes: basic and clinical aspects.Behring Inst Mitt. 1993 Dec;(93):292-8. Behring Inst Mitt. 1993. PMID: 8172579 Review.
Cited by
-
Development of a novel method for analyzing collagen O-glycosylations by hydrazide chemistry.Mol Cell Proteomics. 2012 Jun;11(6):M111.010397. doi: 10.1074/mcp.M111.010397. Epub 2012 Jan 13. Mol Cell Proteomics. 2012. PMID: 22247541 Free PMC article.
-
Mapping surface accessibility of the C1r/C1s tetramer by chemical modification and mass spectrometry provides new insights into assembly of the human C1 complex.J Biol Chem. 2010 Oct 15;285(42):32251-63. doi: 10.1074/jbc.M110.149112. Epub 2010 Jun 30. J Biol Chem. 2010. PMID: 20592021 Free PMC article.
-
Top-down proteomics in health and disease: challenges and opportunities.Proteomics. 2014 May;14(10):1195-210. doi: 10.1002/pmic.201300432. Proteomics. 2014. PMID: 24723472 Free PMC article. Review.
-
C1q: A fresh look upon an old molecule.Mol Immunol. 2017 Sep;89:73-83. doi: 10.1016/j.molimm.2017.05.025. Epub 2017 Jun 7. Mol Immunol. 2017. PMID: 28601358 Free PMC article. Review.
-
A Collagen Triple Helix without the Superhelical Twist.ACS Cent Sci. 2025 Feb 4;11(2):331-345. doi: 10.1021/acscentsci.5c00018. eCollection 2025 Feb 26. ACS Cent Sci. 2025. PMID: 40028357 Free PMC article.
References
-
- Gaboriaud C., Thielens N. M., Gregory L. A., Rossi V., Fontecilla-Camps J. C., Arlaud G. J. (2004) Structure and activation of the C1 complex of complement: unraveling the puzzle. Trends Immunol 25, 368–373 - PubMed
-
- Busby T. F., Ingham K. C. (1990) Amino-terminal calcium-binding domain of human complement C1s mediates the interaction of C1r with C1q. Biochemistry 29, 4613–4618 - PubMed
-
- Illy C., Thielens N. M., Arlaud G. J. (1993) Chemical characterization and location of ionic interactions involved in the assembly of the C1 complex of human complement. J. Protein Chem 12, 771–781 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

