Sex-specific and lineage-specific alternative splicing in primates
- PMID: 20009012
- PMCID: PMC2813474
- DOI: 10.1101/gr.099226.109
Sex-specific and lineage-specific alternative splicing in primates
Abstract
Comparative studies of gene regulation suggest an important role for natural selection in shaping gene expression patterns within and between species. Most of these studies, however, estimated gene expression levels using microarray probes designed to hybridize to only a small proportion of each gene. Here, we used recently developed RNA sequencing protocols, which sidestep this limitation, to assess intra- and interspecies variation in gene regulatory processes in considerably more detail than was previously possible. Specifically, we used RNA-seq to study transcript levels in humans, chimpanzees, and rhesus macaques, using liver RNA samples from three males and three females from each species. Our approach allowed us to identify a large number of genes whose expression levels likely evolve under natural selection in primates. These include a subset of genes with conserved sexually dimorphic expression patterns across the three species, which we found to be enriched for genes involved in lipid metabolism. Our data also suggest that while alternative splicing is tightly regulated within and between species, sex-specific and lineage-specific changes in the expression of different splice forms are also frequent. Intriguingly, among genes in which a change in exon usage occurred exclusively in the human lineage, we found an enrichment of genes involved in anatomical structure and morphogenesis, raising the possibility that differences in the regulation of alternative splicing have been an important force in human evolution.
Figures

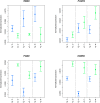
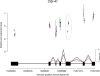
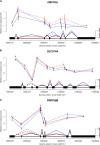
References
-
- Abzhanov A, Protas M, Grant BR, Grant PR, Tabin CJ. Bmp4 and morphological variation of beaks in Darwin's finches. Science. 2004;305:1462–1465. - PubMed
-
- Balashova VA, Abdulkadyrov KM. Cellular composition of hemopoietic tissue of the liver and spleen in the human fetus. Arkh Anat Gistol Embriol. 1984;86:80–83. - PubMed
-
- Britten RJ, Davidson EH. Repetitive and non-repetitive DNA sequences and a speculation on the origins of evolutionary novelty. Q Rev Biol. 1971;46:111–138. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
