Paradox of mistranslation of serine for alanine caused by AlaRS recognition dilemma
- PMID: 20010690
- PMCID: PMC2799227
- DOI: 10.1038/nature08612
Paradox of mistranslation of serine for alanine caused by AlaRS recognition dilemma
Abstract
Mistranslation arising from confusion of serine for alanine by alanyl-tRNA synthetases (AlaRSs) has profound functional consequences. Throughout evolution, two editing checkpoints prevent disease-causing mistranslation from confusing glycine or serine for alanine at the active site of AlaRS. In both bacteria and mice, Ser poses a bigger challenge than Gly. One checkpoint is the AlaRS editing centre, and the other is from widely distributed AlaXps-free-standing, genome-encoded editing proteins that clear Ser-tRNA(Ala). The paradox of misincorporating both a smaller (glycine) and a larger (serine) amino acid suggests a deep conflict for nature-designed AlaRS. Here we show the chemical basis for this conflict. Nine crystal structures, together with kinetic and mutational analysis, provided snapshots of adenylate formation for each amino acid. An inherent dilemma is posed by constraints of a structural design that pins down the alpha-amino group of the bound amino acid by using an acidic residue. This design, dating back more than 3 billion years, creates a serendipitous interaction with the serine OH that is difficult to avoid. Apparently because no better architecture for the recognition of alanine could be found, the serine misactivation problem was solved through free-standing AlaXps, which appeared contemporaneously with early AlaRSs. The results reveal unconventional problems and solutions arising from the historical design of the protein synthesis machinery.
Figures

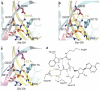


Similar articles
-
Substrate specificity and catalysis by the editing active site of Alanyl-tRNA synthetase from Escherichia coli.Biochemistry. 2011 Mar 8;50(9):1474-82. doi: 10.1021/bi1013535. Epub 2011 Jan 31. Biochemistry. 2011. PMID: 21241052 Free PMC article.
-
Perseverance of protein homeostasis despite mistranslation of glycine codons with alanine.Philos Trans R Soc Lond B Biol Sci. 2023 Feb 27;378(1871):20220029. doi: 10.1098/rstb.2022.0029. Epub 2023 Jan 11. Philos Trans R Soc Lond B Biol Sci. 2023. PMID: 36633285 Free PMC article.
-
The C-Ala domain brings together editing and aminoacylation functions on one tRNA.Science. 2009 Aug 7;325(5941):744-7. doi: 10.1126/science.1174343. Science. 2009. PMID: 19661429 Free PMC article.
-
Alanine transfer RNA synthetase: structure-function relationships and molecular recognition of transfer RNA.Adv Enzymol Relat Areas Mol Biol. 1990;63:233-70. doi: 10.1002/9780470123096.ch4. Adv Enzymol Relat Areas Mol Biol. 1990. PMID: 2407064 Review. No abstract available.
-
The uniqueness of AlaRS and its human disease connections.RNA Biol. 2021 Nov;18(11):1501-1511. doi: 10.1080/15476286.2020.1861803. Epub 2020 Dec 23. RNA Biol. 2021. PMID: 33317386 Free PMC article. Review.
Cited by
-
LOTTE-seq (Long hairpin oligonucleotide based tRNA high-throughput sequencing): specific selection of tRNAs with 3'-CCA end for high-throughput sequencing.RNA Biol. 2020 Jan;17(1):23-32. doi: 10.1080/15476286.2019.1664250. Epub 2019 Sep 16. RNA Biol. 2020. PMID: 31486704 Free PMC article.
-
Emerging roles of lysine lactyltransferases and lactylation.Nat Cell Biol. 2025 Apr;27(4):563-574. doi: 10.1038/s41556-025-01635-8. Epub 2025 Apr 4. Nat Cell Biol. 2025. PMID: 40185947 Review.
-
A molecular link between cell wall biosynthesis, translation fidelity, and stringent response in Streptococcus pneumoniae.Proc Natl Acad Sci U S A. 2021 Apr 6;118(14):e2018089118. doi: 10.1073/pnas.2018089118. Proc Natl Acad Sci U S A. 2021. PMID: 33785594 Free PMC article.
-
Errors in translational decoding: tRNA wobbling or misincorporation?PLoS Genet. 2019 Mar 28;15(3):e1008017. doi: 10.1371/journal.pgen.1008017. eCollection 2019 Mar. PLoS Genet. 2019. PMID: 30921315 Free PMC article. Review.
-
Design principles and functional basis of enantioselectivity of alanyl-tRNA synthetase and a chiral proofreader during protein biosynthesis.Nucleic Acids Res. 2023 Apr 24;51(7):3327-3340. doi: 10.1093/nar/gkad205. Nucleic Acids Res. 2023. PMID: 36951106 Free PMC article.
References
-
- Lee JW, et al. Editing-defective tRNA synthetase causes protein misfolding and neurodegeneration. Nature. 2006;443:50–5. - PubMed
-
- Beebe K, Mock M, Merriman E, Schimmel P. Distinct domains of tRNA synthetase recognize the same base pair. Nature. 2008;451:90–3. - PubMed
-
- Carter CW., Jr. Cognition, mechanism, and evolutionary relationships in aminoacyl-tRNA synthetases. Annu Rev Biochem. 1993;62:715–48. - PubMed
-
- Giege R. The early history of tRNA recognition by aminoacyl-tRNA synthetases. J Biosci. 2006;31:477–88. - PubMed
METHODS REFERENCES
-
- Malde AK, Mark AE. Binding and enantiomeric selectivity of threonyl-tRNA synthetase. J Am Chem Soc. 2009;131:3848–9. - PubMed
-
- Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997;276:307–326. - PubMed
-
- CCP4 The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 1994;50:760–3. - PubMed
-
- Emsley P, Cowtan K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 2004;60:2126–32. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

