RNA-directed DNA methylation and plant development require an IWR1-type transcription factor
- PMID: 20010803
- PMCID: PMC2816620
- DOI: 10.1038/embor.2009.246
RNA-directed DNA methylation and plant development require an IWR1-type transcription factor
Abstract
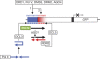
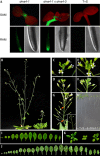
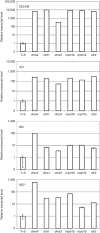
RNA-directed DNA methylation (RdDM) in plants requires two RNA polymerase (Pol) II-related RNA polymerases, namely Pol IV and Pol V. A genetic screen designed to reveal factors that are important for RdDM in a developmental context in Arabidopsis identified DEFECTIVE IN MERISTEM SILENCING 4 (DMS4). Unlike other mutants defective in RdDM, dms4 mutants have a pleiotropic developmental phenotype. The DMS4 protein is similar to yeast IWR1 (interacts with RNA polymerase II), a conserved putative transcription factor that interacts with Pol II subunits. The DMS4 complementary DNA partly complements the K1 killer toxin hypersensitivity of a yeast iwr1 mutant, suggesting some functional conservation. In the transgenic system studied, mutations in DMS4 directly or indirectly affect Pol IV-dependent secondary short interfering RNAs, Pol V-mediated RdDM, Pol V-dependent synthesis of intergenic non-coding RNA and expression of many Pol II-driven genes. These data suggest that DMS4 might be a regulatory factor for several RNA polymerases, thus explaining its diverse roles in the plant.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
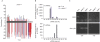
Similar articles
-
NRPD4, a protein related to the RPB4 subunit of RNA polymerase II, is a component of RNA polymerases IV and V and is required for RNA-directed DNA methylation.Genes Dev. 2009 Feb 1;23(3):318-30. doi: 10.1101/gad.1765209. Genes Dev. 2009. PMID: 19204117 Free PMC article.
-
A conserved transcriptional regulator is required for RNA-directed DNA methylation and plant development.Genes Dev. 2009 Dec 1;23(23):2717-22. doi: 10.1101/gad.1851809. Epub 2009 Nov 10. Genes Dev. 2009. PMID: 19903758 Free PMC article.
-
The RNA silencing enzyme RNA polymerase v is required for plant immunity.PLoS Genet. 2011 Dec;7(12):e1002434. doi: 10.1371/journal.pgen.1002434. Epub 2011 Dec 29. PLoS Genet. 2011. PMID: 22242006 Free PMC article.
-
Molecular mechanisms of the RNA polymerases in plant RNA-directed DNA methylation.Trends Biochem Sci. 2024 Mar;49(3):247-256. doi: 10.1016/j.tibs.2023.11.005. Epub 2023 Dec 9. Trends Biochem Sci. 2024. PMID: 38072749 Review.
-
RNA-Directed DNA Methylation: The Evolution of a Complex Epigenetic Pathway in Flowering Plants.Annu Rev Plant Biol. 2015;66:243-67. doi: 10.1146/annurev-arplant-043014-114633. Epub 2014 Dec 10. Annu Rev Plant Biol. 2015. PMID: 25494460 Review.
Cited by
-
A Role for PICKLE in the Regulation of Cold and Salt Stress Tolerance in Arabidopsis.Front Plant Sci. 2019 Jul 9;10:900. doi: 10.3389/fpls.2019.00900. eCollection 2019. Front Plant Sci. 2019. PMID: 31354770 Free PMC article.
-
Highly reproducible label free quantitative proteomic analysis of RNA polymerase complexes.Mol Cell Proteomics. 2011 Feb;10(2):M110.000687. doi: 10.1074/mcp.M110.000687. Epub 2010 Nov 3. Mol Cell Proteomics. 2011. PMID: 21048197 Free PMC article.
-
Identification of genes required for de novo DNA methylation in Arabidopsis.Epigenetics. 2011 Mar;6(3):344-54. doi: 10.4161/epi.6.3.14242. Epub 2011 Mar 1. Epigenetics. 2011. PMID: 21150311 Free PMC article.
-
Regulation and function of DNA methylation in plants and animals.Cell Res. 2011 Mar;21(3):442-65. doi: 10.1038/cr.2011.23. Epub 2011 Feb 15. Cell Res. 2011. PMID: 21321601 Free PMC article. Review.
-
GFP Loss-of-Function Mutations in Arabidopsis thaliana.G3 (Bethesda). 2015 Jul 6;5(9):1849-55. doi: 10.1534/g3.115.019604. G3 (Bethesda). 2015. PMID: 26153075 Free PMC article.
References
-
- Aouida M, Pagé N, Leduc A, Peter M, Ramotar D (2004) A genome-wide screen in Saccharomyces cerevisiae reveals altered transport as a mechanism of resistance to the anticancer drug bleomycin. Cancer Res 64: 1102–1109 - PubMed
-
- Collins S et al. (2007) Functional dissection of protein complexes involved in yeast chromosome biology using a genetic interaction map. Nature 446: 806–810 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

