Meta-analyses of genome-wide association studies identify multiple loci associated with pulmonary function
- PMID: 20010835
- PMCID: PMC2832852
- DOI: 10.1038/ng.500
Meta-analyses of genome-wide association studies identify multiple loci associated with pulmonary function
Abstract
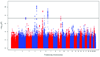
Spirometric measures of lung function are heritable traits that reflect respiratory health and predict morbidity and mortality. We meta-analyzed genome-wide association studies for two clinically important lung-function measures: forced expiratory volume in the first second (FEV(1)) and its ratio to forced vital capacity (FEV(1)/FVC), an indicator of airflow obstruction. This meta-analysis included 20,890 participants of European ancestry from four CHARGE Consortium studies: Atherosclerosis Risk in Communities, Cardiovascular Health Study, Framingham Heart Study and Rotterdam Study. We identified eight loci associated with FEV(1)/FVC (HHIP, GPR126, ADAM19, AGER-PPT2, FAM13A, PTCH1, PID1 and HTR4) and one locus associated with FEV(1) (INTS12-GSTCD-NPNT) at or near genome-wide significance (P < 5 x 10(-8)) in the CHARGE Consortium dataset. Our findings may offer insights into pulmonary function and pathogenesis of chronic lung disease.
Figures










Comment in
-
Lung function and airway diseases.Nat Genet. 2010 Jan;42(1):14-6. doi: 10.1038/ng0110-14. Nat Genet. 2010. PMID: 20037613
References
-
- Wilk JB, et al. Evidence for major genes influencing pulmonary function in the NHLBI family heart study. Genet Epidemiol. 2000;19:81–94. - PubMed
-
- Schunemann HJ, Dorn J, Grant BJ, Winkelstein W, Jr, Trevisan M. Pulmonary function is a long-term predictor of mortality in the general population: 29-year follow-up of the Buffalo Health Study. Chest. 2000;118:656–664. - PubMed
-
- DeMeo DL, Silverman EK. Genetics of chronic obstructive pulmonary disease. Semin Respir Crit Care Med. 2003;24:151–160. - PubMed
-
- Pauwels RA, Buist AS, Calverley PM, Jenkins CR, Hurd SS. Global strategy for the diagnosis, management, and prevention of chronic obstructive pulmonary disease. NHLBI/WHO Global Initiative for Chronic Obstructive Lung Disease (GOLD) Workshop summary. Am J Respir Crit Care Med. 2001;163:1256–1276. - PubMed
Publication types
MeSH terms
Grants and funding
- N01 HC055016/HL/NHLBI NIH HHS/United States
- N01 HC055019/HL/NHLBI NIH HHS/United States
- R01HL59367/HL/NHLBI NIH HHS/United States
- R01 HL077612/HL/NHLBI NIH HHS/United States
- R01 HL086694/HL/NHLBI NIH HHS/United States
- U01HG004402/HG/NHGRI NIH HHS/United States
- N01 HC055020/HL/NHLBI NIH HHS/United States
- DK063491/DK/NIDDK NIH HHS/United States
- N01 HC055022/HL/NHLBI NIH HHS/United States
- ZIA ES043012/ImNIH/Intramural NIH HHS/United States
- R01HL086694/HL/NHLBI NIH HHS/United States
- M01-RR00425/RR/NCRR NIH HHS/United States
- N02-HL-6-4278/HL/NHLBI NIH HHS/United States
- N01-HC-25195/HC/NHLBI NIH HHS/United States
- G0000934/MRC_/Medical Research Council/United Kingdom
- U01 HL080295/HL/NHLBI NIH HHS/United States
- N01-HC-55022/HC/NHLBI NIH HHS/United States
- R01 HL059367/HL/NHLBI NIH HHS/United States
- N01 HC055021/HL/NHLBI NIH HHS/United States
- N01 HC085086/HL/NHLBI NIH HHS/United States
- N01-HC-55016/HC/NHLBI NIH HHS/United States
- N01 HC055015/HL/NHLBI NIH HHS/United States
- RC1 HL100543/HL/NHLBI NIH HHS/United States
- N01-HC-85079 THROUGH N01-HC-85086/HC/NHLBI NIH HHS/United States
- R01 HL087652/HL/NHLBI NIH HHS/United States
- N01-HC-55021/HC/NHLBI NIH HHS/United States
- HHSN268200625226C/PHS HHS/United States
- U01 HG004402/HG/NHGRI NIH HHS/United States
- P30 DK063491/DK/NIDDK NIH HHS/United States
- N01 HC-55222/HC/NHLBI NIH HHS/United States
- N01-HC-55019/HC/NHLBI NIH HHS/United States
- N01 HC025195/HL/NHLBI NIH HHS/United States
- R01HL087641/HL/NHLBI NIH HHS/United States
- N01-HC-55015/HC/NHLBI NIH HHS/United States
- N01 HC055222/HL/NHLBI NIH HHS/United States
- N01 HC085079/HL/NHLBI NIH HHS/United States
- N01-HC-75150/HC/NHLBI NIH HHS/United States
- N01-HC-55020/HC/NHLBI NIH HHS/United States
- N01 HC055018/HL/NHLBI NIH HHS/United States
- Z01 ES043012/ImNIH/Intramural NIH HHS/United States
- M01 RR000425/RR/NCRR NIH HHS/United States
- N01 HC075150/HL/NHLBI NIH HHS/United States
- N01-HC-55018/HC/NHLBI NIH HHS/United States
- R01 HL087641/HL/NHLBI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

