Solution structure and dynamics of human ubiquitin conjugating enzyme Ube2g2
- PMID: 20014027
- PMCID: PMC2822102
- DOI: 10.1002/prot.22648
Solution structure and dynamics of human ubiquitin conjugating enzyme Ube2g2
Abstract
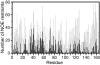
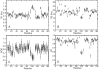
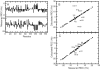
Ube2g2 is an E2 enzyme which functions as part of the endoplasmic reticulum-associated degradation (ERAD) pathway responsible for identification and degradation of misfolded proteins in the endoplasmic reticulum. In tandem with a cognate E3 ligase, Ube2g2 assembles K48-linked polyubiquitin chains and then transfers them to substrate, leading ultimately to proteasomal degradation of the polyubiquitin-tagged substrate. We report here the solution structure and backbone dynamics of Ube2g2 solved by nuclear magnetic resonance spectroscopy. Although the solution structure agrees well with crystallographic structures for the E2 core, catalytically important loops (encompassing residues 95-107 and 130-135) flanking the active site cysteine are poorly defined. (15)N spin relaxation and residual dipolar coupling analysis directly demonstrates that these two loops are highly dynamic in solution. These results suggest that Ube2g2 requires one or more of its protein partners, such as cognate E3, acceptor ubiquitin substrate or thiolester-linked donor ubiquitin, to assume its catalytically relevant conformation. Within the NMR structural ensemble, interactions were observed between His94 and the highly mobile loop residues Asp98 and Asp99, supporting a possible role for His94 as a general base activated by the carboxylate side-chains of Asp98 or Asp99.
Keywords: ERAD; NMR spectroscopy; UBC7; Ube2g2; spin relaxation; ubiquitin conjugating enzyme.
Figures

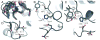
Similar articles
-
Structure of human ubiquitin-conjugating enzyme E2 G2 (UBE2G2/UBC7).Acta Crystallogr Sect F Struct Biol Cryst Commun. 2006 Apr 1;62(Pt 4):330-4. doi: 10.1107/S1744309106009006. Epub 2006 Mar 25. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2006. PMID: 16582478 Free PMC article.
-
Mechanistic insights into active site-associated polyubiquitination by the ubiquitin-conjugating enzyme Ube2g2.Proc Natl Acad Sci U S A. 2009 Mar 10;106(10):3722-7. doi: 10.1073/pnas.0808564106. Epub 2009 Feb 17. Proc Natl Acad Sci U S A. 2009. PMID: 19223579 Free PMC article.
-
A ubiquitin ligase transfers preformed polyubiquitin chains from a conjugating enzyme to a substrate.Nature. 2007 Mar 15;446(7133):333-7. doi: 10.1038/nature05542. Epub 2007 Feb 18. Nature. 2007. PMID: 17310145
-
Dimeric Ube2g2 simultaneously engages donor and acceptor ubiquitins to form Lys48-linked ubiquitin chains.EMBO J. 2014 Jan 7;33(1):46-61. doi: 10.1002/embj.201385315. Epub 2013 Dec 23. EMBO J. 2014. PMID: 24366945 Free PMC article.
-
Diverse roles of the E2/E3 hybrid enzyme UBE2O in the regulation of protein ubiquitination, cellular functions, and disease onset.FEBS J. 2019 Jun;286(11):2018-2034. doi: 10.1111/febs.14708. Epub 2018 Dec 4. FEBS J. 2019. PMID: 30468556 Review.
Cited by
-
Mechanistic insights revealed by a UBE2A mutation linked to intellectual disability.Nat Chem Biol. 2019 Jan;15(1):62-70. doi: 10.1038/s41589-018-0177-2. Epub 2018 Dec 10. Nat Chem Biol. 2019. PMID: 30531907 Free PMC article.
-
Differential ubiquitin binding by the acidic loops of Ube2g1 and Ube2r1 enzymes distinguishes their Lys-48-ubiquitylation activities.J Biol Chem. 2015 Jan 23;290(4):2251-63. doi: 10.1074/jbc.M114.624809. Epub 2014 Dec 3. J Biol Chem. 2015. PMID: 25471371 Free PMC article.
-
An acidic loop and cognate phosphorylation sites define a molecular switch that modulates ubiquitin charging activity in Cdc34-like enzymes.PLoS Comput Biol. 2011 May;7(5):e1002056. doi: 10.1371/journal.pcbi.1002056. Epub 2011 May 26. PLoS Comput Biol. 2011. PMID: 21637798 Free PMC article.
-
Multimodal mechanism of action for the Cdc34 acidic loop: a case study for why ubiquitin-conjugating enzymes have loops and tails.J Biol Chem. 2013 Nov 29;288(48):34882-96. doi: 10.1074/jbc.M113.509190. Epub 2013 Oct 15. J Biol Chem. 2013. PMID: 24129577 Free PMC article.
-
Structure and functional determinants of Rad6-Bre1 subunits in the histone H2B ubiquitin-conjugating complex.Nucleic Acids Res. 2023 Mar 21;51(5):2117-2136. doi: 10.1093/nar/gkad012. Nucleic Acids Res. 2023. PMID: 36715322 Free PMC article.
References
-
- Katsanis N, Fisher EM. Identification, expression, and chromosomal localization of ubiquitin conjugating enzyme 7 (UBE2G2), a human homologue of the Saccharomyces cerevisiae ubc7 gene. Genomics. 1998;51(1):128–131. - PubMed
-
- Kim I, Xu W, Reed JC. Cell death and endoplasmic reticulum stress: disease relevance and therapeutic opportunities. Nat Rev Drug Discov. 2008;7(12):1013–1030. - PubMed
-
- Simoes-Correia J, Figueiredo J, Oliveira C, van Hengel J, Seruca R, van Roy F, Suriano G. Endoplasmic reticulum quality control: a new mechanism of E-cadherin regulation and its implication in cancer. Hum Mol Genet. 2008;17(22):3566–3576. - PubMed
-
- Li W, Tu D, Brunger AT, Ye Y. A ubiquitin ligase transfers preformed polyubiquitin chains from a conjugating enzyme to a substrate. Nature. 2007;446(7133):333–337. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

