Testing the balanced electrostatic interaction hypothesis of hepatitis B virus DNA synthesis by using an in vivo charge rebalance approach
- PMID: 20015989
- PMCID: PMC2820918
- DOI: 10.1128/JVI.01666-09
Testing the balanced electrostatic interaction hypothesis of hepatitis B virus DNA synthesis by using an in vivo charge rebalance approach
Abstract
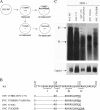
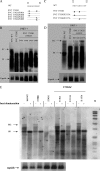
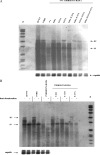
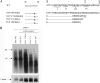
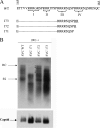
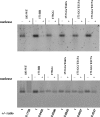
Previously, a charge balance hypothesis was proposed to explain hepatitis B virus (HBV) capsid stability, assembly, RNA encapsidation, and DNA replication. This hypothesis emphasized the importance of a balanced electrostatic interaction between the positive charge from the arginine-rich domain (ARD) of the core protein (HBc) and the negative charge from the encapsidated nucleic acid. It remains unclear if any of the negative charge involved in this electrostatic interaction could come from the HBc protein per se, in addition to the encapsidated nucleic acid. HBc ARD IV mutant 173GG and ARD II mutant 173RR/R157A/R158A are arginine deficient and replication defective. Not surprisingly, the replication defect of ARD IV mutant 173GG can be rescued by restoring positively charged amino acids at the adjacent positions 174 and 175. However, most interestingly, it can be at least partially rescued by reducing negatively charged residues in the assembly domain, such as by glutamic acid-to-alanine (E-to-A) substitutions at position 46 or 117 and to a much lesser extent at position 113. Similar results were obtained for ARD II mutant 173RR/R157A/R158A. These amino acids are located on the inner surfaces of HBc icosahedral particles, and their acidic side chains point toward the capsid interior. For HBV DNA synthesis, the relative amount of positive versus negative charge in the electrostatic interactions is more important than the absolute amount of positive or negative charge. These results support the concept that balanced electrostatic interaction is important during the viral life cycle.
Figures

Similar articles
-
HBV maintains electrostatic homeostasis by modulating negative charges from phosphoserine and encapsidated nucleic acids.Sci Rep. 2016 Dec 13;6:38959. doi: 10.1038/srep38959. Sci Rep. 2016. PMID: 27958343 Free PMC article.
-
Nucleic acid chaperone activity associated with the arginine-rich domain of human hepatitis B virus core protein.J Virol. 2014 Mar;88(5):2530-43. doi: 10.1128/JVI.03235-13. Epub 2013 Dec 18. J Virol. 2014. PMID: 24352445 Free PMC article.
-
Testing an electrostatic interaction hypothesis of hepatitis B virus capsid stability by using an in vitro capsid disassembly/reassembly system.J Virol. 2009 Oct;83(20):10616-26. doi: 10.1128/JVI.00749-09. Epub 2009 Aug 5. J Virol. 2009. PMID: 19656897 Free PMC article.
-
Phosphorylation of the Arginine-Rich C-Terminal Domains of the Hepatitis B Virus (HBV) Core Protein as a Fine Regulator of the Interaction between HBc and Nucleic Acid.Viruses. 2020 Jul 8;12(7):738. doi: 10.3390/v12070738. Viruses. 2020. PMID: 32650547 Free PMC article. Review.
-
Virus Isoelectric Point Estimation: Theories and Methods.Appl Environ Microbiol. 2021 Jan 15;87(3):e02319-20. doi: 10.1128/AEM.02319-20. Print 2021 Jan 15. Appl Environ Microbiol. 2021. PMID: 33188001 Free PMC article. Review.
Cited by
-
Targeting the multifunctional HBV core protein as a potential cure for chronic hepatitis B.Antiviral Res. 2020 Oct;182:104917. doi: 10.1016/j.antiviral.2020.104917. Epub 2020 Aug 17. Antiviral Res. 2020. PMID: 32818519 Free PMC article. Review.
-
Tg1.4HBV-s-rec mice, a crossbred hepatitis B virus-transgenic model, develop mild hepatitis.Sci Rep. 2023 Dec 20;13(1):22829. doi: 10.1038/s41598-023-50090-8. Sci Rep. 2023. PMID: 38129531 Free PMC article.
-
Serine phosphoacceptor sites within the core protein of hepatitis B virus contribute to genome replication pleiotropically.PLoS One. 2011 Feb 15;6(2):e17202. doi: 10.1371/journal.pone.0017202. PLoS One. 2011. PMID: 21358805 Free PMC article.
-
A nuanced role of the small loop of hepatitis B virus small envelope protein in virion morphogenesis and secretion.J Biomed Sci. 2021 Dec 1;28(1):82. doi: 10.1186/s12929-021-00780-0. J Biomed Sci. 2021. PMID: 34852809 Free PMC article.
-
A theoretical model for the dynamic structure of hepatitis B nucleocapsid.Biophys J. 2011 Nov 16;101(10):2476-84. doi: 10.1016/j.bpj.2011.10.002. Epub 2011 Nov 15. Biophys J. 2011. PMID: 22098746 Free PMC article.
References
-
- Beames, B., and R. E. Lanford. 1993. Carboxy-terminal truncations of the HBV core protein affect capsid formation and the apparent size of encapsidated HBV RNA. Virology 194:597-607. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

