Whole genome transcript profiling from fingerstick blood samples: a comparison and feasibility study
- PMID: 20017944
- PMCID: PMC2811129
- DOI: 10.1186/1471-2164-10-617
Whole genome transcript profiling from fingerstick blood samples: a comparison and feasibility study
Abstract
Background: Whole genome gene expression profiling has revolutionized research in the past decade especially with the advent of microarrays. Recently, there have been significant improvements in whole blood RNA isolation techniques which, through stabilization of RNA at the time of sample collection, avoid bias and artifacts introduced during sample handling. Despite these improvements, current human whole blood RNA stabilization/isolation kits are limited by the requirement of a venous blood sample of at least 2.5 mL. While fingerstick blood collection has been used for many different assays, there has yet to be a kit developed to isolate high quality RNA for use in gene expression studies from such small human samples. The clinical and field testing advantages of obtaining reliable and reproducible gene expression data from a fingerstick are many; it is less invasive, time saving, more mobile, and eliminates the need of a trained phlebotomist. Furthermore, this method could also be employed in small animal studies, i.e. mice, where larger sample collections often require sacrificing the animal. In this study, we offer a rapid and simple method to extract sufficient amounts of high quality total RNA from approximately 70 microl of whole blood collected via a fingerstick using a modified protocol of the commercially available Qiagen PAXgene RNA Blood Kit.
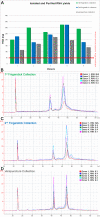
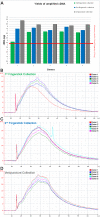
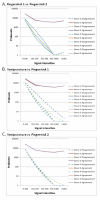
Results: From two sets of fingerstick collections, about 70 uL whole blood collected via finger lancet and capillary tube, we recovered an average of 252.6 ng total RNA with an average RIN of 9.3. The post-amplification yields for 50 ng of total RNA averaged at 7.0 ug cDNA. The cDNA hybridized to Affymetrix HG-U133 Plus 2.0 GeneChips had an average % Present call of 52.5%. Both fingerstick collections were highly correlated with r(2) values ranging from 0.94 to 0.97. Similarly both fingerstick collections were highly correlated to the venous collection with r(2) values ranging from 0.88 to 0.96 for fingerstick collection 1 and 0.94 to 0.96 for fingerstick collection 2.
Conclusions: Our comparisons of RNA quality and gene expression data of the fingerstick method with traditionally processed sample workflows demonstrate excellent RNA quality from the capillary collection as well as very high correlations of gene expression data.
Figures
Similar articles
-
Differential gene expression profiles are dependent upon method of peripheral blood collection and RNA isolation.BMC Genomics. 2008 Oct 10;9:474. doi: 10.1186/1471-2164-9-474. BMC Genomics. 2008. PMID: 18847473 Free PMC article.
-
Gene expression differences between PAXgene and Tempus blood RNA tubes are highly reproducible between independent samples and biobanks.BMC Res Notes. 2017 Mar 23;10(1):136. doi: 10.1186/s13104-017-2455-6. BMC Res Notes. 2017. PMID: 28335817 Free PMC article.
-
Investigating gene expression profiles of whole blood and peripheral blood mononuclear cells using multiple collection and processing methods.PLoS One. 2019 Dec 6;14(12):e0225137. doi: 10.1371/journal.pone.0225137. eCollection 2019. PLoS One. 2019. PMID: 31809517 Free PMC article.
-
RNA stabilization of peripheral blood and profiling by bead chip analysis.Methods Mol Biol. 2009;496:175-210. doi: 10.1007/978-1-59745-553-4_13. Methods Mol Biol. 2009. PMID: 18839112 Review.
-
Genome-wide platelet RNA profiling in clinical samples.Methods Mol Biol. 2009;496:273-83. doi: 10.1007/978-1-59745-553-4_17. Methods Mol Biol. 2009. PMID: 18839116 Review.
Cited by
-
Evaluation of a solid matrix for collection and ambient storage of RNA from whole blood.BMC Clin Pathol. 2014 May 13;14:22. doi: 10.1186/1472-6890-14-22. eCollection 2014. BMC Clin Pathol. 2014. PMID: 24855452 Free PMC article.
-
An Assessment of Individual Preference for a Novel Capillary Blood Collection System.Patient Prefer Adherence. 2024 Mar 1;18:531-541. doi: 10.2147/PPA.S437969. eCollection 2024. Patient Prefer Adherence. 2024. PMID: 38444755 Free PMC article.
-
Single cell profiling of capillary blood enables out of clinic human immunity studies.Sci Rep. 2020 Nov 25;10(1):20540. doi: 10.1038/s41598-020-77073-3. Sci Rep. 2020. PMID: 33239690 Free PMC article.
-
Technical assessment of different extraction methods and transcriptome profiling of RNA isolated from small volumes of blood.Sci Rep. 2023 Mar 3;13(1):3598. doi: 10.1038/s41598-023-30629-5. Sci Rep. 2023. PMID: 36869090 Free PMC article.
-
Blood handling and leukocyte isolation methods impact the global transcriptome of immune cells.BMC Immunol. 2018 Oct 30;19(1):30. doi: 10.1186/s12865-018-0268-6. BMC Immunol. 2018. PMID: 30376808 Free PMC article.
References
-
- Rainen L, Oelmueller U, Jurgensen S, Wyrich R, Ballas C, Schram J, Herdman C, Bankaitis-Davis D, Nicholls N, Trollinger D. Stabilization of mRNA expression in whole blood samples. Clin Chem. 2002;48(11):1883–1890. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

