A role for suppressed incisor cuspal morphogenesis in the evolution of mammalian heterodont dentition
- PMID: 20018657
- PMCID: PMC2806730
- DOI: 10.1073/pnas.0907236107
A role for suppressed incisor cuspal morphogenesis in the evolution of mammalian heterodont dentition
Abstract
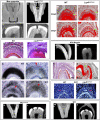
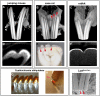
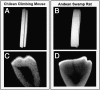
Changes in tooth shape have played a major role in vertebrate evolution with modification of dentition allowing an organism to adapt to new feeding strategies. The current view is that molar teeth evolved from simple conical teeth, similar to canines, by progressive addition of extra "cones" to form progressively complex multicuspid crowns. Mammalian incisors, however, are neither conical nor multicuspid, and their evolution is unclear. We show that hypomorphic mutation of a cell surface receptor, Lrp4, which modulates multiple signaling pathways, produces incisors with grooved enamel surfaces that exhibit the same molecular characteristics as the tips of molar cusps. Mice with a null mutation of Lrp4 develop extra cusps on molars and have incisors that exhibit clear molar-like cusp and root morphologies. Molecular analysis identifies misregulation of Shh and Bmp signaling in the mutant incisors and suggests an uncoupling of the processes of tooth shape determination and morphogenesis. Incisors thus possess a developmentally suppressed, cuspid crown-like morphogenesis program similar to that in molars that is revealed by loss of Lrp4 activity. Several mammalian species naturally possess multicuspid incisors, suggesting that mammals have the capacity to form multicuspid teeth regardless of location in the oral jaw. Localized loss of enamel may thus have been an intermediary step in the evolution of cusps, both of which use Lrp4-mediated signaling.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Morphoregulation of teeth: modulating the number, size, shape and differentiation by tuning Bmp activity.Evol Dev. 2005 Sep-Oct;7(5):440-57. doi: 10.1111/j.1525-142X.2005.05048.x. Evol Dev. 2005. PMID: 16174037 Free PMC article.
-
Lrp4: A novel modulator of extracellular signaling in craniofacial organogenesis.Am J Med Genet A. 2010 Dec;152A(12):2974-83. doi: 10.1002/ajmg.a.33372. Am J Med Genet A. 2010. PMID: 21108386 Free PMC article. Review.
-
Tooth morphogenesis and ameloblast differentiation are regulated by micro-RNAs.Dev Biol. 2010 Apr 15;340(2):355-68. doi: 10.1016/j.ydbio.2010.01.019. Epub 2010 Jan 25. Dev Biol. 2010. PMID: 20102707
-
Enamel knots as signaling centers linking tooth morphogenesis and odontoblast differentiation.Adv Dent Res. 2001 Aug;15:14-8. doi: 10.1177/08959374010150010401. Adv Dent Res. 2001. PMID: 12640732 Review.
-
Lrp4 modulates extracellular integration of cell signaling pathways in development.PLoS One. 2008;3(12):e4092. doi: 10.1371/journal.pone.0004092. Epub 2008 Dec 31. PLoS One. 2008. PMID: 19116665 Free PMC article.
Cited by
-
Rare Variants in LRP4 Are Associated with Mesiodens, Root Maldevelopment, and Oral Exostoses in Humans.Biology (Basel). 2023 Jan 30;12(2):220. doi: 10.3390/biology12020220. Biology (Basel). 2023. PMID: 36829498 Free PMC article.
-
Common developmental pathways link tooth shape to regeneration.Dev Biol. 2013 May 15;377(2):399-414. doi: 10.1016/j.ydbio.2013.02.007. Epub 2013 Feb 17. Dev Biol. 2013. PMID: 23422830 Free PMC article.
-
RSK2 is a modulator of craniofacial development.PLoS One. 2014 Jan 8;9(1):e84343. doi: 10.1371/journal.pone.0084343. eCollection 2014. PLoS One. 2014. PMID: 24416220 Free PMC article.
-
Development and evolution of the unique cetacean dentition.PeerJ. 2013 Feb 19;1:e24. doi: 10.7717/peerj.24. Print 2013. PeerJ. 2013. PMID: 23638359 Free PMC article.
-
Neutron scanning reveals unexpected complexity in the enamel thickness of an herbivorous Jurassic reptile.J R Soc Interface. 2018 Jun;15(143):20180039. doi: 10.1098/rsif.2018.0039. J R Soc Interface. 2018. PMID: 29899156 Free PMC article.
References
-
- Rose KD. The Beginning of the Age of Mammals. Baltimore: Johns Hopkins University Press; 2006.
-
- Tummers M, Yamashiro T, Thesleff I. Modulation of epithelial cell fate of the root in vitro. J Dent Res. 2007;86:1063–1067. - PubMed
-
- Tummers M, Thesleff I. Observations on continuously growing roots of the sloth and the K14-Eda transgenic mice indicate that epithelial stem cells can give rise to both the ameloblast and root epithelium cell lineage creating distinct tooth patterns. Evol Dev. 2008;10:187–195. - PubMed
-
- Nykjaer A, Willnow TE. The low-density lipoprotein receptor gene family: A cellular Swiss army knife? Trends Cell Biol. 2002;12:273–280. - PubMed
-
- Herz J, Bock HH. Lipoprotein receptors in the nervous system. Annu Rev Biochem. 2002;71:405–434. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

