Probing ribosome-nascent chain complexes produced in vivo by NMR spectroscopy
- PMID: 20018739
- PMCID: PMC2799734
- DOI: 10.1073/pnas.0903750106
Probing ribosome-nascent chain complexes produced in vivo by NMR spectroscopy
Abstract
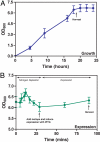
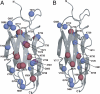
The means by which a polypeptide chain acquires its unique 3-D structure is a fundamental question in biology. During its synthesis on the ribosome, a nascent chain (NC) emerges vectorially and will begin to fold in a cotranslational fashion. The complex environment of the cell, coupled with the gradual emergence of the ribosome-tethered NC during its synthesis, imposes conformational restraints on its folding landscape that differ from those placed on an isolated protein when stimulated to fold following denaturation in solution. To begin to examine cotranslational folding as it would occur within a cell, we produce highly selective, isotopically labeled NCs bound to isotopically silent ribosomes in vivo. We then apply NMR spectroscopy to study, at a residue specific level, the conformation of NCs consisting of different fractional lengths of the polypeptide chain corresponding to a given protein. This combined approach provides a powerful means of generating a series of snapshots of the folding of the NC as it emerges from the ribosome. Application of this strategy to the NMR analysis of the progressive synthesis of an Ig-like domain reveals the existence of a partially folded ribosome-bound species that is likely to represent an intermediate species populated during the cotranslational folding process.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

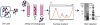


Similar articles
-
A structural ensemble of a ribosome-nascent chain complex during cotranslational protein folding.Nat Struct Mol Biol. 2016 Apr;23(4):278-285. doi: 10.1038/nsmb.3182. Epub 2016 Feb 29. Nat Struct Mol Biol. 2016. PMID: 26926436 Free PMC article.
-
Structure and dynamics of a ribosome-bound nascent chain by NMR spectroscopy.Proc Natl Acad Sci U S A. 2007 Oct 16;104(42):16516-21. doi: 10.1073/pnas.0704664104. Epub 2007 Oct 10. Proc Natl Acad Sci U S A. 2007. PMID: 17940046 Free PMC article.
-
A strategy for co-translational folding studies of ribosome-bound nascent chain complexes using NMR spectroscopy.Nat Protoc. 2016 Aug;11(8):1492-507. doi: 10.1038/nprot.2016.101. Epub 2016 Jul 28. Nat Protoc. 2016. PMID: 27466710
-
The ribosome and its role in protein folding: looking through a magnifying glass.Acta Crystallogr D Struct Biol. 2017 Jun 1;73(Pt 6):509-521. doi: 10.1107/S2059798317007446. Epub 2017 May 31. Acta Crystallogr D Struct Biol. 2017. PMID: 28580913 Free PMC article. Review.
-
Cotranslational Folding of Proteins on the Ribosome.Biomolecules. 2020 Jan 7;10(1):97. doi: 10.3390/biom10010097. Biomolecules. 2020. PMID: 31936054 Free PMC article. Review.
Cited by
-
Solid-state NMR enhanced by dynamic nuclear polarization as a novel tool for ribosome structural biology.J Biomol NMR. 2013 Jun;56(2):85-93. doi: 10.1007/s10858-013-9721-2. Epub 2013 May 21. J Biomol NMR. 2013. PMID: 23689811
-
Structural polymorphism and substrate promiscuity of a ribosome-associated molecular chaperone.Magn Reson (Gott). 2021 Jun 4;2(1):375-386. doi: 10.5194/mr-2-375-2021. eCollection 2021. Magn Reson (Gott). 2021. PMID: 37904759 Free PMC article.
-
Conformational flexibility within the nascent polypeptide-associated complex enables its interactions with structurally diverse client proteins.J Biol Chem. 2018 Jun 1;293(22):8554-8568. doi: 10.1074/jbc.RA117.001568. Epub 2018 Apr 12. J Biol Chem. 2018. PMID: 29650757 Free PMC article.
-
Mechanically unfolding protein L using a laser-feedback-controlled cantilever.Biophys J. 2011 Apr 6;100(7):1800-9. doi: 10.1016/j.bpj.2011.02.021. Biophys J. 2011. PMID: 21463594 Free PMC article.
-
Nascent chains derived from a foldable protein sequence interact with specific ribosomal surface sites near the exit tunnel.Sci Rep. 2024 May 29;14(1):12324. doi: 10.1038/s41598-024-61274-1. Sci Rep. 2024. PMID: 38811604 Free PMC article.
References
-
- Steitz TA. A structural understanding of the dynamic ribosome machine. Nat Rev Mol Cell Biol. 2008;9:242–253. - PubMed
-
- Bashan A, Yonath A. Correlating ribosome function with high-resolution structures. Trends Microbiol. 2008;16:326–335. - PubMed
-
- Korostelev A, Trakhanov S, Laurberg M, Noller HF. Crystal structure of a 70S ribosome-tRNA complex reveals functional interactions and rearrangements. Cell. 2006;126:1065–1077. - PubMed
-
- Mitra K, Frank J. Ribosome dynamics: Insights from atomic structure modeling into cryo-electron microscopy maps. Ann Rev Biophys Biomol Struc. 2006;35:299–317. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

