Fetal hemoglobin in sickle cell anemia: genome-wide association studies suggest a regulatory region in the 5' olfactory receptor gene cluster
- PMID: 20018918
- PMCID: PMC2832816
- DOI: 10.1182/blood-2009-08-239517
Fetal hemoglobin in sickle cell anemia: genome-wide association studies suggest a regulatory region in the 5' olfactory receptor gene cluster
Abstract
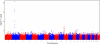
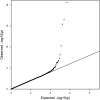
In a genome-wide association study of 848 blacks with sickle cell anemia, we identified single nucleotide polymorphisms (SNPs) associated with fetal hemoglobin concentration. The most significant SNPs in a discovery sample were tested in a replication set of 305 blacks with sickle cell anemia and in subjects with hemoglobin E or beta thalassemia trait from Thailand and Hong Kong. A novel region on chromosome 11 containing olfactory receptor genes OR51B5 and OR51B6 was identified by 6 SNPs (lowest P = 4.7E-08) and validated in the replication set. An additional olfactory receptor gene, OR51B2, was identified by a novel SNP set enrichment analysis. Genome-wide association studies also validated a previously identified SNP (rs766432) in BCL11A, a gene known to affect fetal hemoglobin levels (P = 2.6E-21) and in Thailand and Hong Kong subjects. Elements within the olfactory receptor gene cluster might play a regulatory role in gamma-globin gene expression.
Figures

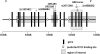
Comment in
-
Extensive admixture in Brazilian sickle cell patients: implications for the mapping of genetic modifiers.Blood. 2011 Oct 20;118(16):4493-5; author reply 4495. doi: 10.1182/blood-2011-06-361915. Blood. 2011. PMID: 22021456 Free PMC article. No abstract available.
References
-
- Steinberg MH, Nagel RL. Genetic modulation of sickle cell disease and thalassemia. In: Steinberg MH, Forget BG, Higgs DR, Weatherall DJ, editors. Disorders of Hemoglobin: Genetics, Pathophysiology and Clinical Management. Cambridge, United Kingdom: Cambridge University Press; 2009. pp. 638–657.
-
- Garner C, Tatu T, Reittie JE, et al. Genetic influences on F cells and other hematologic variables: a twin heritability study. Blood. 2000;95(1):342–346. - PubMed
-
- Menzel S, Thein SL. Genetic architecture of hemoglobin F control. Curr Opin Hematol. 2009;16(3):179–186. - PubMed
-
- Gaston M, Rosse WF. The cooperative study of sickle cell disease: review of study design and objectives. Am J Pediatr Hematol Oncol. 1982;4(2):197–201. - PubMed
-
- Steinberg MH, Lu ZH, Barton FB, Terrin ML, Charache S, Dover GJ. Fetal hemoglobin in sickle cell anemia: determinants of response to hydroxyurea. Multicenter Study of Hydroxyurea. Blood. 1997;89(3):1078–1088. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

