Algorithms for locating extremely conserved elements in multiple sequence alignments
- PMID: 20021665
- PMCID: PMC2808710
- DOI: 10.1186/1471-2105-10-432
Algorithms for locating extremely conserved elements in multiple sequence alignments
Abstract
Background: In 2004, Bejerano et al. announced the startling discovery of hundreds of "ultraconserved elements", long genomic sequences perfectly conserved across human, mouse, and rat. Their announcement stimulated a flurry of subsequent research.
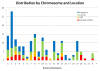
Results: We generalize the notion of ultraconserved element in a natural way from extraordinary human-rodent conservation to extraordinary conservation over an arbitrary set of species. We call these "Extremely Conserved Elements". There is a linear time algorithm to find all such Extremely Conserved Elements in any multiple sequence alignment, provided that the conservation is required to be across all the aligned species. For the general case of conservation across an arbitrary subset of the aligned species, we show that the question of whether there exists an Extremely Conserved Element is NP-complete. We illustrate the linear time algorithm by cataloguing all 177 Extremely Conserved Elements in the currently available 44-vertebrate whole-genome alignment, and point out some of the characteristics of these elements.
Conclusions: The NP-completeness in the case of conservation across an arbitrary subset of the aligned species implies that it is unlikely an efficient algorithm exists for this general case. Despite this fact, for the interesting case of conservation across all or most of the aligned species, our algorithm is efficient enough to be practical. The 177 Extremely Conserved Elements that we catalog demonstrate many of the characteristics of the original ultraconserved elements of Bejerano et al.
Figures
References
-
- Sakuraba Y, Kimura T, Masuya H, Noguchi H, Sezutsu H, Takahasi KR, Toyoda A, Fukumura R, Murata T, Sakaki Y, Yamamura M, Wakana S, Noda T, Shiroishi T, Gondo Y. Identification and characterization of new long conserved noncoding sequences in vertebrates. Mammalian Genome. 2008;19:703–712. doi: 10.1007/s00335-008-9152-7. - DOI - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous