tRNA gene sequences are required for transcriptional silencing in Entamoeba histolytica
- PMID: 20023072
- PMCID: PMC2823001
- DOI: 10.1128/EC.00248-09
tRNA gene sequences are required for transcriptional silencing in Entamoeba histolytica
Abstract
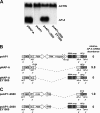
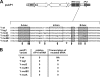
Transcriptional silencing by trans inactivation can contribute to the regulation of gene expression in eukaryotic cells. In the human intestinal protozoan parasite Entamoeba histolytica, trans inactivation of the amoebapore-A gene (AP-A) was recently achieved by episomal transfection of E. histolytica trophozoites with the plasmid psAP1. The mechanism of AP-A trans inactivation is largely unknown, though it was suggested that a partial short interspersed transposable element (SINE) is required. By systematic assessment of various E. histolytica isolates transfected with psAP1 derivates, trans inactivation of AP-A was restricted to the strain HM-1:IMSS (2411) but could not be achieved in other standard laboratory strains. Importantly, sequences of an E. histolytica tRNA array that were located on psAP1 in close proximity to the AP-A upstream region and comprising the glutamic acid (TTC) (E) and tyrosine (GTA) (Y) tRNA genes were indispensable for AP-A silencing. In contrast to the case described in previous reports, SINE was not required for AP-A trans inactivation. AP-A expression could be regained in silenced cells by episomal transfection under the control of a heterologous E. histolytica promoter, opening a way toward future silencing of individual genes of interest in E. histolytica. Our results indicate that tRNA gene-mediated silencing is not restricted to Saccharomyces cerevisiae.
Figures




References
-
- Andrulis E. D., Neiman A. M., Zappulla D. C., Sternglanz R. 1998. Perinuclear localization of chromatin facilitates transcriptional silencing. Nature 394:592–595 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

