Comparison of PCR binary typing (P-BIT), a new approach to epidemiological subtyping of Campylobacter jejuni, with serotyping, pulsed-field gel electrophoresis, and multilocus sequence typing methods
- PMID: 20023103
- PMCID: PMC2832355
- DOI: 10.1128/AEM.02215-09
Comparison of PCR binary typing (P-BIT), a new approach to epidemiological subtyping of Campylobacter jejuni, with serotyping, pulsed-field gel electrophoresis, and multilocus sequence typing methods
Abstract
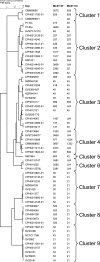
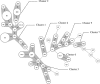
To overcome some of the deficiencies with current molecular typing schema for Campylobacter spp., we developed a prototype PCR binary typing (P-BIT) approach. We investigated the distribution of 68 gene targets in 58 Campylobacter jejuni strains, one Campylobacter lari strain, and two Campylobacter coli strains for this purpose. Gene targets were selected on the basis of distribution in multiple genomes or plasmids, and known or putative status as an epidemicity factor. Strains were examined with Penner serotyping, pulsed-field gel electrophoresis (PFGE; using SmaI and KpnI enzymes), and multilocus sequence typing (MLST) approaches for comparison. P-BIT provided 100% typeability for strains and gave a diversity index of 98.5%, compared with 97.0% for SmaI PFGE, 99.4% for KpnI PFGE, 96.1% for MLST, and 92.8% for serotyping. Numerical analysis of the P-BIT data clearly distinguished strains of the three Campylobacter species examined and correlated somewhat with MLST clonal complex assignations and with previous classifications of "high" and "low" risk. We identified 18 gene targets that conferred the same level of discrimination as the 68 initially examined. We conclude that P-BIT is a useful approach for subtyping, offering advantages of speed, cost, and potential for strain risk ranking unavailable from current molecular typing schema for Campylobacter spp.
Figures

Similar articles
-
Designing multiplex PCR system of Campylobacter jejuni for efficient typing by improving monoplex PCR binary typing method.J Infect Chemother. 2015 Jan;21(1):50-4. doi: 10.1016/j.jiac.2014.09.003. Epub 2014 Nov 5. J Infect Chemother. 2015. PMID: 25455748
-
Utility of multilocus sequence typing as an epidemiological tool for investigation of outbreaks of gastroenteritis caused by Campylobacter jejuni.J Clin Microbiol. 2003 Oct;41(10):4733-9. doi: 10.1128/JCM.41.10.4733-4739.2003. J Clin Microbiol. 2003. PMID: 14532212 Free PMC article.
-
Typing of Campylobacter jejuni isolates from dogs by use of multilocus sequence typing and pulsed-field gel electrophoresis.J Clin Microbiol. 2009 Nov;47(11):3466-71. doi: 10.1128/JCM.01046-09. Epub 2009 Sep 30. J Clin Microbiol. 2009. PMID: 19794053 Free PMC article.
-
Phenotypic and genotypic methods for typing Campylobacter jejuni and Campylobacter coli in poultry.Poult Sci. 2012 Jan;91(1):255-64. doi: 10.3382/ps.2011-01414. Poult Sci. 2012. PMID: 22184452 Review.
-
A systematic review and meta-analysis of Penner serotype prevalence of Campylobacter jejuni in low- and middle-income countries.PLoS One. 2021 May 5;16(5):e0251039. doi: 10.1371/journal.pone.0251039. eCollection 2021. PLoS One. 2021. PMID: 33951106 Free PMC article.
Cited by
-
Aerotolerance and Multi-Locus Sequence Typing of Campylobacter jejuni Isolated from Commercial Broiler Processing Plants.Foods. 2023 Sep 2;12(17):3305. doi: 10.3390/foods12173305. Foods. 2023. PMID: 37685237 Free PMC article.
-
Molecular Typing of Campylobacter jejuni and Campylobacter coli Isolated from Various Retail Meats by MLST and PFGE.Foods. 2014 Jan 8;3(1):82-93. doi: 10.3390/foods3010082. Foods. 2014. PMID: 28234305 Free PMC article.
-
Development of a DNA-based microarray for the detection of zoonotic pathogens in rodent species.Mol Cell Probes. 2015 Dec;29(6):427-437. doi: 10.1016/j.mcp.2015.07.005. Epub 2015 Jul 15. Mol Cell Probes. 2015. PMID: 26188129 Free PMC article.
-
Comprehensive detection and discrimination of Campylobacter species by use of confocal micro-Raman spectroscopy and multilocus sequence typing.J Clin Microbiol. 2012 Sep;50(9):2932-46. doi: 10.1128/JCM.01144-12. Epub 2012 Jun 27. J Clin Microbiol. 2012. PMID: 22740711 Free PMC article.
-
Development and validation of a comparative genomic fingerprinting method for high-resolution genotyping of Campylobacter jejuni.J Clin Microbiol. 2012 Mar;50(3):788-97. doi: 10.1128/JCM.00669-11. Epub 2011 Dec 14. J Clin Microbiol. 2012. PMID: 22170908 Free PMC article.
References
-
- Bereswill, S., and M. Kist. 2003. Recent developments in Campylobacter pathogenesis. Curr. Opin. Infect. Dis. 16:487-491. - PubMed
-
- Centers for Disease Control and Prevention. 2009. Campylobacter jejuni infection associated with unpasteurized milk and cheese—Kansas, 2007. MMWR Morb. Mortal. Wkly. Rep. 57:1377-1379. - PubMed
-
- Champion, O. L., M. W. Gaunt, O. Gundogdu, A. Elmi, A. A. Witney, J. Hinds, N. Dorrell, and B. W. Wren. 2005. Comparative phylogenomics of the food-borne pathogen Campylobacter jejuni reveals genetic markers predictive of infection source. Proc. Natl. Acad. Sci. U. S. A. 102:16043-16048. - PMC - PubMed
-
- Chou, W. K., S. Dick, W. W. Wakarchuk, and M. E. Tanner. 2005. Identification and characterization of NeuB3 from Campylobacter jejuni as a pseudaminic acid synthase. J. Biol. Chem. 280:35922-35928. - PubMed
-
- Devane, M. L., C. Nicol, A. Ball, J. D. Klena, P. Scholes, J. A. Hudson, M. G. Baker, B. J. Gilpin, N. Garrett, and M. G. Savill. 2005. The occurrence of Campylobacter subtypes in environmental reservoirs and potential transmission routes. J. Appl. Microbiol. 98:980-990. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

