Borrelia burgdorferi locus BB0795 encodes a BamA orthologue required for growth and efficient localization of outer membrane proteins
- PMID: 20025662
- PMCID: PMC5581936
- DOI: 10.1111/j.1365-2958.2009.07015.x
Borrelia burgdorferi locus BB0795 encodes a BamA orthologue required for growth and efficient localization of outer membrane proteins
Abstract
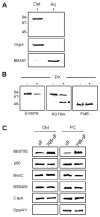
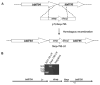
The outer membrane (OM) of the pathogenic diderm spirochete, Borrelia burgdorferi, contains integral beta-barrel outer membrane proteins (OMPs) in addition to its numerous outer surface lipoproteins. Very few OMPs have been identified in B. burgdorferi, and the protein machinery required for OMP assembly and OM localization is currently unknown. Essential OM BamA proteins have recently been characterized in Gram-negative bacteria that are central components of an OM beta-barrel assembly machine and are required for proper localization and insertion of bacterial OMPs. In the present study, we characterized a putative B. burgdorferi BamA orthologue encoded by open reading frame bb0795. Structural model predictions and cellular localization data indicate that the B. burgdorferi BB0795 protein contains an N-terminal periplasmic domain and a C-terminal, surface-exposed beta-barrel domain. Additionally, assays with an IPTG-regulatable bb0795 mutant revealed that BB0795 is required for B. burgdorferi growth. Furthermore, depletion of BB0795 results in decreased amounts of detectable OMPs in the B. burgdorferi OM. Interestingly, a decrease in the levels of surface-exposed lipoproteins was also observed in the mutant OMs. Collectively, our structural, cellular localization and functional data are consistent with the characteristics of other BamA proteins, indicating that BB0795 is a B. burgdorferi BamA orthologue.
Figures
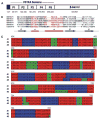


 ), 0.05 mM (
), 0.05 mM (
 ) or 1 mM (
) or 1 mM (
 ) IPTG in triplicate. The parental B31-A3-LK strain (
) IPTG in triplicate. The parental B31-A3-LK strain (
 ) was seeded in IPTG-deplete media. Spirochetes were enumerated daily by dark-field microscopy for 13 days post inoculation. B. IPTG dose-dependent expression of BB0795 in B. burgdorferi strain flacp-795-LK. Whole-cell lysates of B. burgdorferi flacp-795-LK cultivated in media containing 0.05 mM or 1 mM IPTG were subjected to immunoblot analysis using rat anti-BB0795 antibodies. Lysates were also probed with a LacI monoclonal antibody and FlaB antisera to ensure equivalent amounts of the LacI repressor protein and equivalent amounts of whole-cell lysates were loaded, respectively. Immunoblots of whole-cell lysates of the parental B31-A3-LK strain (grown in IPTG-deplete media) are shown for comparison.
) was seeded in IPTG-deplete media. Spirochetes were enumerated daily by dark-field microscopy for 13 days post inoculation. B. IPTG dose-dependent expression of BB0795 in B. burgdorferi strain flacp-795-LK. Whole-cell lysates of B. burgdorferi flacp-795-LK cultivated in media containing 0.05 mM or 1 mM IPTG were subjected to immunoblot analysis using rat anti-BB0795 antibodies. Lysates were also probed with a LacI monoclonal antibody and FlaB antisera to ensure equivalent amounts of the LacI repressor protein and equivalent amounts of whole-cell lysates were loaded, respectively. Immunoblots of whole-cell lysates of the parental B31-A3-LK strain (grown in IPTG-deplete media) are shown for comparison.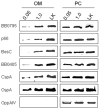
Similar articles
-
Characterization of the β-barrel assembly machine accessory lipoproteins from Borrelia burgdorferi.BMC Microbiol. 2015 Mar 24;15:70. doi: 10.1186/s12866-015-0411-y. BMC Microbiol. 2015. PMID: 25887384 Free PMC article.
-
BB0324 and BB0028 are constituents of the Borrelia burgdorferi β-barrel assembly machine (BAM) complex.BMC Microbiol. 2012 Apr 20;12:60. doi: 10.1186/1471-2180-12-60. BMC Microbiol. 2012. PMID: 22519960 Free PMC article.
-
A conserved C-terminal domain of TamB interacts with multiple BamA POTRA domains in Borreliella burgdorferi.PLoS One. 2024 Aug 29;19(8):e0304839. doi: 10.1371/journal.pone.0304839. eCollection 2024. PLoS One. 2024. PMID: 39208212 Free PMC article.
-
The β-Barrel Assembly Machinery Complex.Methods Mol Biol. 2015;1329:1-16. doi: 10.1007/978-1-4939-2871-2_1. Methods Mol Biol. 2015. PMID: 26427672 Review.
-
The Treponema pallidum Outer Membrane.Curr Top Microbiol Immunol. 2018;415:1-38. doi: 10.1007/82_2017_44. Curr Top Microbiol Immunol. 2018. PMID: 28849315 Free PMC article. Review.
Cited by
-
Probing the Borrelia burgdorferi surface lipoprotein secretion pathway using a conditionally folding protein domain.J Bacteriol. 2011 Dec;193(23):6724-32. doi: 10.1128/JB.06042-11. Epub 2011 Sep 30. J Bacteriol. 2011. PMID: 21965569 Free PMC article.
-
Analyzing the molecular mechanism of lipoprotein localization in Brucella.Front Microbiol. 2015 Oct 28;6:1189. doi: 10.3389/fmicb.2015.01189. eCollection 2015. Front Microbiol. 2015. PMID: 26579096 Free PMC article. Review.
-
Outer membrane lipoproteins: late to the party, but the center of attention.J Bacteriol. 2025 Jan 31;207(1):e0044224. doi: 10.1128/jb.00442-24. Epub 2024 Dec 13. J Bacteriol. 2025. PMID: 39670753 Free PMC article. Review.
-
Borrelia burgdorferi Surface Exposed GroEL Is a Multifunctional Protein.Pathogens. 2021 Feb 18;10(2):226. doi: 10.3390/pathogens10020226. Pathogens. 2021. PMID: 33670728 Free PMC article.
-
BamE modulates the Escherichia coli beta-barrel assembly machine component BamA.J Bacteriol. 2012 Mar;194(5):1002-8. doi: 10.1128/JB.06426-11. Epub 2011 Dec 16. J Bacteriol. 2012. PMID: 22178970 Free PMC article.
References
-
- Arnold K, Bordoli L, Kopp J, Schwede T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics. 2006;22:195–201. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

