Quality assessment of DNA derived from up to 30 years old formalin fixed paraffin embedded (FFPE) tissue for PCR-based methylation analysis using SMART-MSP and MS-HRM
- PMID: 20025721
- PMCID: PMC2804714
- DOI: 10.1186/1471-2407-9-453
Quality assessment of DNA derived from up to 30 years old formalin fixed paraffin embedded (FFPE) tissue for PCR-based methylation analysis using SMART-MSP and MS-HRM
Abstract
Background: The High Resolution Melting (HRM) technology has recently been introduced as a rapid and robust analysis tool for the detection of DNA methylation. The methylation status of multiple tumor suppressor genes may serve as biomarkers for early cancer diagnostics, for prediction of prognosis and for prediction of response to treatment. Therefore, it is important that methodologies for detection of DNA methylation continue to evolve. Sensitive Melting Analysis after Real Time - Methylation Specific PCR (SMART-MSP) and Methylation Sensitive - High Resolution Melting (MS-HRM) are two methods for single locus DNA methylation detection based on HRM.
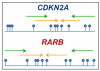
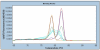
Methods: Here, we have assessed the quality of DNA extracted from up to 30 years old Formalin Fixed Paraffin Embedded (FFPE) tissue for DNA methylation analysis using SMART-MSP and MS-HRM. The quality assessment was performed on DNA extracted from 54 Non-Small Cell Lung Cancer (NSCLC) samples derived from FFPE tissue, collected over 30 years and grouped into five years intervals. For each sample, the methylation levels of the CDKN2A (p16) and RARB promoters were estimated using SMART-MSP and MS-HRM assays designed to assess the methylation status of the same CpG positions. This allowed for a direct comparison of the methylation levels estimated by the two methods for each sample.
Results: CDKN2A promoter methylation levels were successfully determined by SMART-MSP and MS-HRM in all 54 samples. Identical methylation estimates were obtained by the two methods in 46 of the samples. The methylation levels of the RARB promoter were successfully determined by SMART-MSP in all samples. When using MS-HRM to assess RARB methylation five samples failed to amplify and 15 samples showed a melting profile characteristic for heterogeneous methylation. Twenty-seven of the remaining 34 samples, for which the methylation level could be estimated, gave the same result as observed when using SMART-MSP.
Conclusion: MS-HRM and SMART-MSP can be successfully used for single locus methylation studies using DNA derived from up to 30 years old FFPE tissue. Furthermore, it can be expected that MS-HRM and SMART-MSP will provide similar methylation estimates when assays are designed to analyze the same CpG positions.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

