Fusion of short telomeres in human cells is characterized by extensive deletion and microhomology, and can result in complex rearrangements
- PMID: 20026586
- PMCID: PMC2847243
- DOI: 10.1093/nar/gkp1183
Fusion of short telomeres in human cells is characterized by extensive deletion and microhomology, and can result in complex rearrangements
Abstract
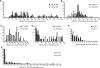
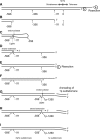
Telomere fusion is an important mutational event that has the potential to lead to large-scale genomic rearrangements of the types frequently observed in cancer. We have developed single-molecule approaches to detect, isolate and characterize the DNA sequence of telomere fusion events in human cells. Using these assays, we have detected complex fusion events that include fusion with interstitial loci adjacent to fragile sites, intra-molecular rearrangements, and fusion events involving the telomeres of both arms of the same chromosome consistent with ring chromosome formation. All fusion events were characterized by the deletion of at least one of the telomeres extending into the sub-telomeric DNA up to 5.6 kb; close to the limit of our assays. The deletion profile indicates that deletion may extend further into the chromosome. Short patches of DNA sequence homology with a G:C bias were observed at the fusion point in 60% of events. The distinct profile that accompanies telomere fusion may be a characteristic of the end-joining processes involved in the fusion event.
Figures



Similar articles
-
Mre11 modulates the fidelity of fusion between short telomeres in human cells.Nucleic Acids Res. 2012 Mar;40(6):2518-26. doi: 10.1093/nar/gkr1117. Epub 2011 Dec 1. Nucleic Acids Res. 2012. PMID: 22139912 Free PMC article.
-
The nature of telomere fusion and a definition of the critical telomere length in human cells.Genes Dev. 2007 Oct 1;21(19):2495-508. doi: 10.1101/gad.439107. Genes Dev. 2007. PMID: 17908935 Free PMC article.
-
Telomere dysfunction and fusion during the progression of chronic lymphocytic leukemia: evidence for a telomere crisis.Blood. 2010 Sep 16;116(11):1899-907. doi: 10.1182/blood-2010-02-272104. Epub 2010 Jun 10. Blood. 2010. PMID: 20538793
-
Telomeres and mechanisms of Robertsonian fusion.Chromosoma. 1998 May;107(2):136-40. doi: 10.1007/s004120050289. Chromosoma. 1998. PMID: 9601982 Review.
-
Telomeres, chromosome instability and cancer.Nucleic Acids Res. 2006 May 8;34(8):2408-17. doi: 10.1093/nar/gkl303. Print 2006. Nucleic Acids Res. 2006. PMID: 16682448 Free PMC article. Review.
Cited by
-
The Response to DNA Damage at Telomeric Repeats and Its Consequences for Telomere Function.Genes (Basel). 2019 Apr 24;10(4):318. doi: 10.3390/genes10040318. Genes (Basel). 2019. PMID: 31022960 Free PMC article. Review.
-
Effect of DNA repair inhibitor AsiDNA on the incidence of telomere fusion in crisis.Hum Mol Genet. 2021 Apr 26;30(3-4):172-181. doi: 10.1093/hmg/ddab008. Hum Mol Genet. 2021. PMID: 33480989 Free PMC article.
-
Telomere fusions associate with coding sequence and copy number alterations in CLL.Leukemia. 2019 Aug;33(8):2093-2097. doi: 10.1038/s41375-019-0423-y. Epub 2019 Feb 22. Leukemia. 2019. PMID: 30796307 Free PMC article. No abstract available.
-
Mre11 modulates the fidelity of fusion between short telomeres in human cells.Nucleic Acids Res. 2012 Mar;40(6):2518-26. doi: 10.1093/nar/gkr1117. Epub 2011 Dec 1. Nucleic Acids Res. 2012. PMID: 22139912 Free PMC article.
-
Telomere length measurement-caveats and a critical assessment of the available technologies and tools.Mutat Res. 2012 Feb 1;730(1-2):59-67. doi: 10.1016/j.mrfmmm.2011.04.003. Epub 2011 Jun 12. Mutat Res. 2012. PMID: 21663926 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

