NPR1 protein regulates pathogenic and symbiotic interactions between Rhizobium and legumes and non-legumes
- PMID: 20027302
- PMCID: PMC2793007
- DOI: 10.1371/journal.pone.0008399
NPR1 protein regulates pathogenic and symbiotic interactions between Rhizobium and legumes and non-legumes
Abstract
Background: Legumes are unique in their ability to establish symbiotic interaction with rhizobacteria from Rhizobium genus, which provide them with available nitrogen. Nodulation factors (NFs) produced by Rhizobium initiate legume root hair deformation and curling that entrap the bacteria, and allow it to grow inside the plant. In contrast, legumes and non-legumes activate defense responses when inoculated with pathogenic bacteria. One major defense pathway is mediated by salicylic acid (SA). SA is sensed and transduced to downstream defense components by a redox-regulated protein called NPR1.
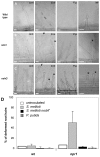
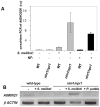
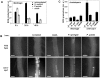
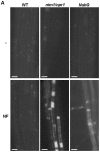
Methodology/principal findings: We used Arabidopsis mutants in SA defense pathway to test the role of NPR1 in symbiotic interactions. Inoculation of Sinorhizobium meliloti or purified NF on Medicago truncatula or nim1/npr1 A. thaliana mutants induced root hair deformation and transcription of early and late nodulins. Application of S. meliloti or NF on M. truncatula or A. thaliana roots also induced a strong oxidative burst that lasted much longer than in plants inoculated with pathogenic or mutualistic bacteria. Transient overexpression of NPR1 in M. truncatula suppressed root hair curling, while inhibition of NPR1 expression by RNAi accelerated curling.
Conclusions/significance: We show that, while NPR1 has a positive effect on pathogen resistance, it has a negative effect on symbiotic interactions, by inhibiting root hair deformation and nodulin expression. Our results also show that basic plant responses to Rhizobium inoculation are conserved in legumes and non-legumes.
Conflict of interest statement
Figures




Similar articles
-
The non-specific lipid transfer protein N5 of Medicago truncatula is implicated in epidermal stages of rhizobium-host interaction.BMC Plant Biol. 2012 Dec 7;12:233. doi: 10.1186/1471-2229-12-233. BMC Plant Biol. 2012. PMID: 23217154 Free PMC article.
-
Root hair curling and Rhizobium infection in Medicago truncatula are mediated by phosphatidylinositide-regulated endocytosis and reactive oxygen species.J Exp Bot. 2007;58(7):1637-49. doi: 10.1093/jxb/erm013. Epub 2007 Apr 9. J Exp Bot. 2007. PMID: 17420174
-
ROS production during symbiotic infection suppresses pathogenesis-related gene expression.Plant Signal Behav. 2012 Mar;7(3):409-15. doi: 10.4161/psb.19217. Epub 2012 Mar 1. Plant Signal Behav. 2012. PMID: 22499208 Free PMC article.
-
Medicago truncatula proteomics.J Proteomics. 2010 Sep 10;73(10):1974-85. doi: 10.1016/j.jprot.2010.07.004. Epub 2010 Jul 16. J Proteomics. 2010. PMID: 20621211 Review.
-
Rhizobial measures to evade host defense strategies and endogenous threats to persistent symbiotic nitrogen fixation: a focus on two legume-rhizobium model systems.Cell Mol Life Sci. 2011 Apr;68(8):1327-39. doi: 10.1007/s00018-011-0650-5. Epub 2011 Mar 2. Cell Mol Life Sci. 2011. PMID: 21365276 Free PMC article. Review.
Cited by
-
Dynamics of Hydrogen Peroxide Accumulation During Tip Growth of Infection Thread in Nodules and Cell Differentiation in Pea (Pisum sativum L.) Symbiotic Nodules.Plants (Basel). 2024 Oct 18;13(20):2923. doi: 10.3390/plants13202923. Plants (Basel). 2024. PMID: 39458872 Free PMC article.
-
Evidence of a role for foliar salicylic acid in regulating the rate of post-ingestive protein breakdown in ruminants and contributing to landscape pollution.J Exp Bot. 2012 May;63(8):3243-55. doi: 10.1093/jxb/ers048. Epub 2012 Feb 29. J Exp Bot. 2012. PMID: 22378947 Free PMC article.
-
Interaction of Medicago truncatula lysin motif receptor-like kinases, NFP and LYK3, produced in Nicotiana benthamiana induces defence-like responses.PLoS One. 2013 Jun 4;8(6):e65055. doi: 10.1371/journal.pone.0065055. Print 2013. PLoS One. 2013. PMID: 23750228 Free PMC article.
-
Oak root response to ectomycorrhizal symbiosis establishment: RNA-Seq derived transcript identification and expression profiling.PLoS One. 2014 May 23;9(5):e98376. doi: 10.1371/journal.pone.0098376. eCollection 2014. PLoS One. 2014. PMID: 24859293 Free PMC article.
-
Reactive Oxygen Species and Nitric Oxide Control Early Steps of the Legume - Rhizobium Symbiotic Interaction.Front Plant Sci. 2016 Apr 8;7:454. doi: 10.3389/fpls.2016.00454. eCollection 2016. Front Plant Sci. 2016. PMID: 27092165 Free PMC article. Review.
References
-
- Schultze M, Kondorosi A. Regulation of symbiotic root nodule development. An Rev Genet. 1998;32:33–57. - PubMed
-
- Durrant WE, Dong X. Systemic acquired resistance. Annual Review of Phytopathology. 2004;42:185–209. - PubMed
-
- Nimchuk Z, Eulgem T, Holt BF, III, Dangl JL. Recognition and response in the plant immune system. Ann Rev Genet. 2003;37:579–609. - PubMed
-
- Mou Z, Fan W, Dong X. Inducers of Plant Systemic Acquired Resistance Regulate NPR1 Function through Redox Changes. Cell. 2003;113:935–944. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

