Unraveling the structure and function of G protein-coupled receptors through NMR spectroscopy
- PMID: 20028318
- PMCID: PMC2801883
- DOI: 10.2174/138161209789824803
Unraveling the structure and function of G protein-coupled receptors through NMR spectroscopy
Abstract
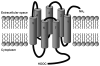
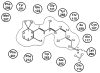
G protein-coupled receptors (GPCRs) are a large superfamily of signaling proteins expressed on the plasma membrane. They are involved in a wide range of physiological processes and, therefore, are exploited as drug targets in a multitude of therapeutic areas. In this extent, knowledge of structural and functional properties of GPCRs may greatly facilitate rational design of modulator compounds. Solution and solid-state nuclear magnetic resonance (NMR) spectroscopy represents a powerful method to gather atomistic insights into protein structure and dynamics. In spite of the difficulties inherent the solution of the structure of membrane proteins through NMR, these methods have been successfully applied, sometimes in combination with molecular modeling, to the determination of the structure of GPCR fragments, the mapping of receptor-ligand interactions, and the study of the conformational changes associated with the activation of the receptors. In this review, we provide a summary of the NMR contributions to the study of the structure and function of GPCRs, also in light of the published crystal structures.
Figures
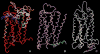
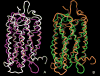
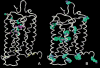
Similar articles
-
G-protein-coupled receptor structure, ligand binding and activation as studied by solid-state NMR spectroscopy.Biochem J. 2013 Mar 15;450(3):443-57. doi: 10.1042/BJ20121644. Biochem J. 2013. PMID: 23445222 Review.
-
Dynamic G Protein-Coupled Receptor Signaling Probed by Solution NMR Spectroscopy.Biochemistry. 2020 Mar 17;59(10):1065-1080. doi: 10.1021/acs.biochem.0c00032. Epub 2020 Mar 2. Biochemistry. 2020. PMID: 32092261
-
Solution- and solid-state NMR studies of GPCRs and their ligands.Biochim Biophys Acta. 2011 Jun;1808(6):1462-75. doi: 10.1016/j.bbamem.2010.10.003. Epub 2010 Oct 15. Biochim Biophys Acta. 2011. PMID: 20951674 Review.
-
Structural biology of human GPCR drugs and endogenous ligands - insights from NMR spectroscopy.Methods. 2020 Aug 1;180:79-88. doi: 10.1016/j.ymeth.2020.08.008. Epub 2020 Sep 8. Methods. 2020. PMID: 32911074 Free PMC article. Review.
-
GPCR drug discovery: integrating solution NMR data with crystal and cryo-EM structures.Nat Rev Drug Discov. 2019 Jan;18(1):59-82. doi: 10.1038/nrd.2018.180. Epub 2018 Nov 9. Nat Rev Drug Discov. 2019. PMID: 30410121 Free PMC article. Review.
Cited by
-
Biophysical characterization of G-protein coupled receptor-peptide ligand binding.Biochem Cell Biol. 2011 Apr;89(2):98-105. doi: 10.1139/o10-142. Biochem Cell Biol. 2011. PMID: 21455262 Free PMC article. Review.
-
Uniform isotope labeling of a eukaryotic seven-transmembrane helical protein in yeast enables high-resolution solid-state NMR studies in the lipid environment.J Biomol NMR. 2011 Feb;49(2):151-61. doi: 10.1007/s10858-011-9473-9. Epub 2011 Jan 19. J Biomol NMR. 2011. PMID: 21246256
-
Small expression tags enhance bacterial expression of the first three transmembrane segments of the apelin receptor.Biochem Cell Biol. 2014 Aug;92(4):269-78. doi: 10.1139/bcb-2014-0009. Epub 2014 May 23. Biochem Cell Biol. 2014. PMID: 24943103 Free PMC article.
-
Mass spectrometry-based proteomics of human cannabinoid receptor 2: covalent cysteine 6.47(257)-ligand interaction affording megagonist receptor activation.J Proteome Res. 2011 Oct 7;10(10):4789-98. doi: 10.1021/pr2005583. Epub 2011 Sep 13. J Proteome Res. 2011. PMID: 21861534 Free PMC article.
-
G protein-coupled receptors involved in GnRH regulation: molecular insights from human disease.Mol Cell Endocrinol. 2011 Oct 22;346(1-2):91-101. doi: 10.1016/j.mce.2011.06.022. Epub 2011 Jun 29. Mol Cell Endocrinol. 2011. PMID: 21736917 Free PMC article. Review.
References
-
- Pierce KL, Premont RT, Lefkowitz RJ. Seven-transmembrane receptors. Nature Reviews Molecular Cell Biology. 2002;3:639–650. - PubMed
-
- Takeda S, Kadowaki S, Haga T, Takaesu H, Mitaku S. Identification of G protein-coupled receptor genes from the human genome sequence. Febs Letters. 2002;520:97–101. - PubMed
-
- Yeagle PL, Albert AD. G-protein coupled receptor structure. Biochimica et Biophysica Acta-Biomembranes. 2007;1768:808–824. - PubMed
-
- Li J, Edwards PC, Burghammer M, Villa C, Schertler GF. Structure of bovine rhodopsin in a trigonal crystal form. J Mol Biol. 2004;343:1409–1438. - PubMed
-
- Okada T, Sugihara M, Bondar AN, Elstner M, Entel P, Buss V. The retinal conformation and its environment in rhodopsin in light of a new 2.2 Å crystal structure. J Mol Biol. 2004;342:571–583. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

