Single strand conformation polymorphism based SNP and Indel markers for genetic mapping and synteny analysis of common bean (Phaseolus vulgaris L.)
- PMID: 20030833
- PMCID: PMC2806352
- DOI: 10.1186/1471-2164-10-629
Single strand conformation polymorphism based SNP and Indel markers for genetic mapping and synteny analysis of common bean (Phaseolus vulgaris L.)
Abstract
Background: Expressed sequence tags (ESTs) are an important source of gene-based markers such as those based on insertion-deletions (Indels) or single-nucleotide polymorphisms (SNPs). Several gel based methods have been reported for the detection of sequence variants, however they have not been widely exploited in common bean, an important legume crop of the developing world. The objectives of this project were to develop and map EST based markers using analysis of single strand conformation polymorphisms (SSCPs), to create a transcript map for common bean and to compare synteny of the common bean map with sequenced chromosomes of other legumes.
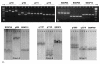
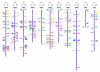
Results: A set of 418 EST based amplicons were evaluated for parental polymorphisms using the SSCP technique and 26% of these presented a clear conformational or size polymorphism between Andean and Mesoamerican genotypes. The amplicon based markers were then used for genetic mapping with segregation analysis performed in the DOR364 x G19833 recombinant inbred line (RIL) population. A total of 118 new marker loci were placed into an integrated molecular map for common bean consisting of 288 markers. Of these, 218 were used for synteny analysis and 186 presented homology with segments of the soybean genome with an e-value lower than 7 x 10-12. The synteny analysis with soybean showed a mosaic pattern of syntenic blocks with most segments of any one common bean linkage group associated with two soybean chromosomes. The analysis with Medicago truncatula and Lotus japonicus presented fewer syntenic regions consistent with the more distant phylogenetic relationship between the galegoid and phaseoloid legumes.
Conclusion: The SSCP technique is a useful and inexpensive alternative to other SNP or Indel detection techniques for saturating the common bean genetic map with functional markers that may be useful in marker assisted selection. In addition, the genetic markers based on ESTs allowed the construction of a transcript map and given their high conservation between species allowed synteny comparisons to be made to sequenced genomes. This synteny analysis may support positional cloning of target genes in common bean through the use of genomic information from these other legumes.
Figures



References
-
- Wang DG, Fan J-B, Siao C-J, Berno A, Young P, Sapolsky R, Ghandour G, Perkins N, Winchester E, Spencer J, Kruglyak L, Stein L, Hsie L, Topaloglou T, Hubbell E, Robinson E, Mittmann M, Morris MS, Shen N, Kilburn D, Rioux J, Nusbaum C, Rozen S, Hudson TJ, Lipshutz R, Chee M, Lander ES. Large-scale identification, mapping, and genotyping of single-nucleotide polymorphisms in the human genome. Science. 1998;280(5366):1077–1082. doi: 10.1126/science.280.5366.1077. - DOI - PubMed
-
- Salmaso M, Faes G, Segala C, Stefanini M, Salakhutdinov I, Zyprian E, Toepfer R, Grando MS, Velasco R. Genome diversity and gene haplotypes in the grapevine (Vitis vinifera L.), as revealed by single nucleotide polymorphisms. Molecular Breeding. 2004;14(4):385–395. doi: 10.1007/s11032-004-0261-z. - DOI
-
- Jones E, Chu W-C, Ayele M, Ho J, Bruggeman E, Yourstone K, Rafalski A, Smith O, McMullen M, Bezawada C, Warren J, Babayev J, Basu S, Smith S. Development of single nucleotide polymorphism (SNP) markers for use in commercial maize (Zea mays L.) germplasm. Molecular Breeding. 2009;24(2):165–176. doi: 10.1007/s11032-009-9281-z. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

