The nonpolymorphic MHC Qa-1b mediates CD8+ T cell surveillance of antigen-processing defects
- PMID: 20038604
- PMCID: PMC2812552
- DOI: 10.1084/jem.20091429
The nonpolymorphic MHC Qa-1b mediates CD8+ T cell surveillance of antigen-processing defects
Erratum in
- J Exp Med. 2010 Mar 15;207(3):671. Drijfhout, Jan W [added]
Abstract
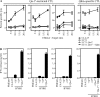
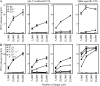
The nonclassical major histocompatibility complex (MHC) Qa-1b accommodates monomorphic leader peptides and functions as a ligand for germ line receptors CD94/NKG2, which are expressed by natural killer cells and CD8+ T cells. We here describe that the conserved peptides are replaced by a novel peptide repertoire of surprising diversity as a result of impairments in the antigen-processing pathway. This novel peptide repertoire represents immunogenic neoantigens for CD8+ T cells, as we found that these Qa-1b-restricted T cells dominantly participated in the response to tumors with processing deficiencies. A surprisingly wide spectrum of target cells, irrespective of transformation status, MHC background, or type of processing deficiency, was recognized by this T cell subset, complying with the conserved nature of Qa-1b. Target cell recognition depended on T cell receptor and Qa-1b interaction, and immunization with identified peptide epitopes demonstrated in vivo priming of CD8+ T cells. Our data reveal that Qa-1b, and most likely its human homologue human leukocyte antigen-E, is important for the defense against processing-deficient cells by displacing the monomorphic leader peptides, which relieves the inhibition through CD94/NKG2A on lymphocytes, and by presenting a novel repertoire of immunogenic peptides, which recruits a subset of cytotoxic CD8+ T cells.
Figures




References
-
- Bijker M.S., van den Eeden S.J., Franken K.L., Melief C.J., Offringa R., van der Burg S.H. 2007. CD8+ CTL priming by exact peptide epitopes in incomplete Freund’s adjuvant induces a vanishing CTL response, whereas long peptides induce sustained CTL reactivity. J. Immunol. 179:5033–5040 - PubMed
-
- Bland F.A., Lemberg M.K., McMichael A.J., Martoglio B., Braud V.M. 2003. Requirement of the proteasome for the trimming of signal peptide-derived epitopes presented by the nonclassical major histocompatibility complex class I molecule HLA-E. J. Biol. Chem. 278:33747–33752 doi: 10.1074/jbc.M305593200 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

