Efavirenz binding to HIV-1 reverse transcriptase monomers and dimers
- PMID: 20039714
- PMCID: PMC2896556
- DOI: 10.1021/bi901579y
Efavirenz binding to HIV-1 reverse transcriptase monomers and dimers
Abstract
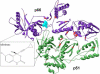
Efavirenz (EFV) is a nonnucleoside reverse transcriptase inhibitor (NNRTI) of HIV-1 reverse transcriptase (RT) used for the treatment of AIDS. RT is a heterodimer composed of p66 and p51 subunits; p51 is produced from p66 by C-terminal truncation by HIV protease. The monomers can form p66/p66 and p51/p51 homodimers as well as the p66/p51 heterodimer. Dimerization and efavirenz binding are coupled processes. In the crystal structure of the p66/p51-EFV complex, the drug is bound to the p66 subunit. The binding of efavirenz to wild-type and dimerization-defective RT proteins was studied by equilibrium dialysis, tryptophan fluorescence, and native gel electrophoresis. A 1:1 binding stoichiometry was determined for both monomers and homodimers. Equilibrium dissociation constants are approximately 2.5 microM for both p66- and p51-EFV complexes, 250 nM for the p66/p66-EFV complex, and 7 nM for the p51/p51-EFV complex. An equilibrium dissociation constant of 92 nM for the p66/p51-EFV complex was calculated from the thermodynamic linkage between dimerization and inhibitor binding. Binding and unbinding kinetics monitored by fluorescence were slow. Progress curve analyses revealed a one-step, direct binding mechanism with association rate constants k(1) of approximately 13.5 M(-1) s(-1) for monomers and heterodimer and dissociation rate constants k(-1) of approximately 9 x 10(-5) s(-1) for monomers. A conformational selection mechanism is proposed to account for the slow association rate. These results show that efavirenz is a slow, tight-binding inhibitor capable of binding all forms of RT and suggest that the NNRTI binding site in monomers and dimers is similar.
Figures


 ) p51 monitored by tryptophan fluorescence at 5 °C. λex = 295 nm, λem = 340 nm. 20 μM p66/p51 (83% heterodimer) diluted to 2 μM prior to adding efavirenz; 4.5 μM p51 (97% monomer). [EFV]:[protein] 2:1.
) p51 monitored by tryptophan fluorescence at 5 °C. λex = 295 nm, λem = 340 nm. 20 μM p66/p51 (83% heterodimer) diluted to 2 μM prior to adding efavirenz; 4.5 μM p51 (97% monomer). [EFV]:[protein] 2:1.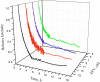
 ) 5 μM, (
) 5 μM, ( ) 8 μM, (
) 8 μM, ( ) 11 μM, and (—) 14 μM efavirenz. Data acquired at 30 s intervals for first 4–5 h, then at 5 min intervals. Data were fit to eq 2a to obtain kobs.
) 11 μM, and (—) 14 μM efavirenz. Data acquired at 30 s intervals for first 4–5 h, then at 5 min intervals. Data were fit to eq 2a to obtain kobs.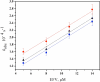
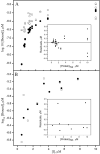
 ) p51W401A, (
) p51W401A, ( ) p66W401A. Lines are fits to eq 3. Error bars give the range of average values from 2 experiments.
) p66W401A. Lines are fits to eq 3. Error bars give the range of average values from 2 experiments.
 ) p51W401A—EFV and (
) p51W401A—EFV and ( ) p66W401A—EFV complexes monitored by tryptophan fluorescence at 5 °C. λex = 295 nm, λem = 340 nm. Black curves are fits to eq 2b. Data acquired at 30 s intervals for first 4–5 h, then at 5 min intervals.
) p66W401A—EFV complexes monitored by tryptophan fluorescence at 5 °C. λex = 295 nm, λem = 340 nm. Black curves are fits to eq 2b. Data acquired at 30 s intervals for first 4–5 h, then at 5 min intervals.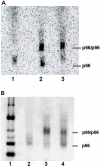
Similar articles
-
Efavirenz binding site in HIV-1 reverse transcriptase monomers.Biochemistry. 2010 Dec 14;49(49):10565-73. doi: 10.1021/bi101480z. Epub 2010 Nov 19. Biochemistry. 2010. PMID: 21090588 Free PMC article.
-
Effects of efavirenz binding on the subunit equilibria of HIV-1 reverse transcriptase.Biochemistry. 2006 Mar 7;45(9):2779-89. doi: 10.1021/bi051915z. Biochemistry. 2006. PMID: 16503633
-
Human immunodeficiency virus type 1 reverse transcriptase dimer destabilization by 1-[Spiro[4"-amino-2",2" -dioxo-1",2" -oxathiole-5",3'-[2', 5'-bis-O-(tert-butyldimethylsilyl)-beta-D-ribofuranosyl]]]-3-ethylthy mine.Biochemistry. 2000 Feb 15;39(6):1427-33. doi: 10.1021/bi991682+. Biochemistry. 2000. PMID: 10684624
-
Efavirenz a nonnucleoside reverse transcriptase inhibitor of first-generation: Approaches based on its medicinal chemistry.Eur J Med Chem. 2016 Jan 27;108:455-465. doi: 10.1016/j.ejmech.2015.11.025. Epub 2015 Nov 25. Eur J Med Chem. 2016. PMID: 26708112 Review.
-
Conformational changes in HIV-1 reverse transcriptase induced by nonnucleoside reverse transcriptase inhibitor binding.Curr HIV Res. 2004 Oct;2(4):323-32. doi: 10.2174/1570162043351093. Curr HIV Res. 2004. PMID: 15544453 Free PMC article. Review.
Cited by
-
The interplay of structure and dynamics: insights from a survey of HIV-1 reverse transcriptase crystal structures.Proteins. 2013 Oct;81(10):1792-801. doi: 10.1002/prot.24325. Epub 2013 Aug 16. Proteins. 2013. PMID: 23720322 Free PMC article.
-
NMR structure of the HIV-1 reverse transcriptase thumb subdomain.J Biomol NMR. 2016 Dec;66(4):273-280. doi: 10.1007/s10858-016-0077-2. Epub 2016 Nov 17. J Biomol NMR. 2016. PMID: 27858311 Free PMC article.
-
Sequence Diversity and Encoded Enzymatic Differences of Monocistronic L1 ORF2 mRNA Variants in the Aged Normal and Alzheimer's Disease Brain.J Neurosci. 2025 Jun 18;45(25):e2298242025. doi: 10.1523/JNEUROSCI.2298-24.2025. J Neurosci. 2025. PMID: 40368603 Free PMC article.
-
Selective killing of human immunodeficiency virus infected cells by non-nucleoside reverse transcriptase inhibitor-induced activation of HIV protease.Retrovirology. 2010 Oct 15;7:89. doi: 10.1186/1742-4690-7-89. Retrovirology. 2010. PMID: 20950436 Free PMC article.
-
Efavirenz binding site in HIV-1 reverse transcriptase monomers.Biochemistry. 2010 Dec 14;49(49):10565-73. doi: 10.1021/bi101480z. Epub 2010 Nov 19. Biochemistry. 2010. PMID: 21090588 Free PMC article.
References
-
- di Marzo Veronese F, Copeland TD, DeVico AL, Rahman R, Oroszlan S, Gallo RC, Sarngadharan MG. Characterization of highly immunogenic p66/p51 as the reverse transcriptase of HTLV-III/LAV. Science. 1986;231:1289–1291. - PubMed
-
- Restle T, Muller B, Goody RS. Dimerization of human immunodeficiency virus type 1 reverse transcriptase. J. Biol. Chem. 1990;265:8986–8988. - PubMed
-
- Restle T, Muller B, Goody RS. RNase H activity of HIV reverse transcriptase is confined exclusively to the dimeric forms. FEBS Lett. 1992;300:97–100. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

