Genome-wide dissection of globally emergent multi-drug resistant serotype 19A Streptococcus pneumoniae
- PMID: 20042094
- PMCID: PMC2807444
- DOI: 10.1186/1471-2164-10-642
Genome-wide dissection of globally emergent multi-drug resistant serotype 19A Streptococcus pneumoniae
Abstract
Background: Emergence of multi-drug resistant (MDR) serotype 19A Streptococcus pneumoniae (SPN) is well-documented but causal factors remain unclear. Canadian SPN isolates (1993-2008, n = 11,083) were serotyped and in vitro susceptibility tested. A subset of MDR 19A were multi-locus sequence typed (MLST) and representative isolates' whole genomes sequenced.
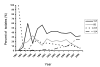
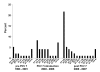
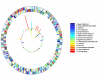
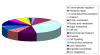
Results: MDR 19A increased in the post-PCV7 era while 19F, 6B, and 23F concurrently declined. MLST of MDR 19A (n = 97) revealed that sequence type (ST) 320 predominated. ST320 was unique amongst MDR 19A in that its minimum inhibitory concentration (MIC) values for penicillin, amoxicillin, ceftriaxone, and erythromycin were higher than for other ST present amongst post-PCV7 MDR 19A. DNA sequencing revealed that alleles at key drug resistance loci pbp2a, pbp2x, pbp2b, ermB, mefA/E, and tetM were conserved between pre-PCV7 ST 320 19F and post-PCV7 ST 320 19A most likely due to a capsule switch recombination event. A genome wide comparison of MDR 19A ST320 with MDR 19F ST320 identified 822 unique SNPs in 19A, 61 of which were present in antimicrobial resistance genes and 100 in virulence factors.
Conclusions: Our results suggest a complex genetic picture where high-level drug resistance, vaccine selection pressure, and SPN mutational events have created a "perfect storm" for the emergence of MDR 19A.
Figures



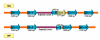
References
-
- Fenoll A, Granizo JJ, Aguilar L, Gimenez MJ, Aragoneses-Fenoll L, Hanquet G, Casal J, Tarrago D. Temporal trends of invasive Streptococcus pneumoniae serotypes and antimicrobial resistance patterns in Spain from 1979 to 2007. J Clin Microbiol. 2009;47:1012–1020. doi: 10.1128/JCM.01454-08. - DOI - PMC - PubMed
-
- Mahjoub-Messai F, Doit C, Koeck JL, Billard T, Evrard B, Bidet P, Hubans C, Raymond J, Levy C, Cohen R, Bingen E. Population snapshot of Streptococcus pneumoniae serotype 19A isolates before and after introduction of seven-valent pneumococcal Vaccination for French children. J Clin Microbiol. 2009;47:837–840. doi: 10.1128/JCM.01547-08. - DOI - PMC - PubMed
-
- Ciccotelli WA, Poutanen SM, Alqahtani M, Morris SK, Cox P, Low DE, Pillai DR, Opavsky MA. A new twist on an old problem. A case of pediatric meningitis caused by multidrug-resistant Streptococcus pneumoniae serotype 19a. Pediatr Infect Dis J. 2009;28:74–75. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

