Proteasome subunits relocalize during human cytomegalovirus infection, and proteasome activity is necessary for efficient viral gene transcription
- PMID: 20042513
- PMCID: PMC2826056
- DOI: 10.1128/JVI.02236-09
Proteasome subunits relocalize during human cytomegalovirus infection, and proteasome activity is necessary for efficient viral gene transcription
Abstract
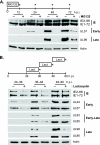
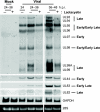
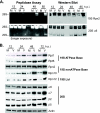
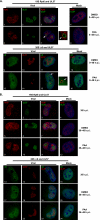
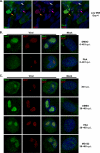
We have continued studies to further understand the role of the ubiquitin-proteasome system (UPS) in human cytomegalovirus (HCMV) infection. With specific inhibitors of the proteasome, we show that ongoing proteasome activity is necessary for facilitating the various stages of the infection. Immediate-early protein 2 expression is modestly reduced with addition of proteasome inhibitors at the onset of infection; however, both early and late gene expression are significantly delayed, even if the inhibitor is removed at 12 h postinfection. Adding the inhibitor at later times during the infection blocks the further accumulation of viral early and late gene products, the severity of which is dependent on when the proteasome is inhibited. This can be attributed primarily to a block in viral RNA transcription, although DNA synthesis is also partially inhibited. Proteasome activity and expression increase as the infection progresses, and this coincides with the relocalization of active proteasomes to the periphery of the viral DNA replication center, where there is active RNA transcription. Interestingly, one 19S subunit, Rpn2, is specifically recruited into the viral DNA replication center. The relocalization of the subunits requires viral DNA replication, but their maintenance around or within the replication center is not dependent on continued viral DNA synthesis or the proteolytic activity of the proteasome. These studies highlight the importance of the UPS at all stages of the HCMV infection and support further studies into this pathway as a potential antiviral target.
Figures





Similar articles
-
The 19S proteasome activator promotes human cytomegalovirus immediate early gene expression through proteolytic and nonproteolytic mechanisms.J Virol. 2014 Oct;88(20):11782-90. doi: 10.1128/JVI.01720-14. Epub 2014 Jul 30. J Virol. 2014. PMID: 25078702 Free PMC article.
-
Proteasome inhibitors: a novel tool to suppress human cytomegalovirus replication and virus-induced immune modulation.Antivir Ther. 2003 Dec;8(6):555-67. Antivir Ther. 2003. PMID: 14760889
-
Human cytomegalovirus IE2 drives transcription initiation from a select subset of late infection viral promoters by host RNA polymerase II.PLoS Pathog. 2020 Apr 6;16(4):e1008402. doi: 10.1371/journal.ppat.1008402. eCollection 2020 Apr. PLoS Pathog. 2020. PMID: 32251483 Free PMC article.
-
Coordination of late gene transcription of human cytomegalovirus with viral DNA synthesis: recombinant viruses as potential therapeutic vaccine candidates.Expert Opin Ther Targets. 2013 Feb;17(2):157-66. doi: 10.1517/14728222.2013.740460. Epub 2012 Dec 12. Expert Opin Ther Targets. 2013. PMID: 23231449 Review.
-
Human cytomegalovirus riding the cell cycle.Med Microbiol Immunol. 2015 Jun;204(3):409-19. doi: 10.1007/s00430-015-0396-z. Epub 2015 Mar 17. Med Microbiol Immunol. 2015. PMID: 25776080 Review.
Cited by
-
Human cytomegalovirus IE2 86 and IE2 40 proteins differentially regulate UL84 protein expression posttranscriptionally in the absence of other viral gene products.J Virol. 2010 May;84(10):5158-70. doi: 10.1128/JVI.00090-10. Epub 2010 Mar 3. J Virol. 2010. PMID: 20200242 Free PMC article.
-
MG132 exerts anti-viral activity against HSV-1 by overcoming virus-mediated suppression of the ERK signaling pathway.Sci Rep. 2020 Apr 21;10(1):6671. doi: 10.1038/s41598-020-63438-1. Sci Rep. 2020. PMID: 32317666 Free PMC article.
-
The ubiquitin-proteasome system is necessary for the replication of duck Tembusu virus.Microb Pathog. 2019 Jul;132:362-368. doi: 10.1016/j.micpath.2019.04.044. Epub 2019 May 1. Microb Pathog. 2019. PMID: 31054366 Free PMC article.
-
Human cytomegalovirus UL76 elicits novel aggresome formation via interaction with S5a of the ubiquitin proteasome system.J Virol. 2013 Nov;87(21):11562-78. doi: 10.1128/JVI.01568-13. Epub 2013 Aug 21. J Virol. 2013. PMID: 23966401 Free PMC article.
-
Viral takeover of the host ubiquitin system.Front Microbiol. 2011 Jul 28;2:161. doi: 10.3389/fmicb.2011.00161. eCollection 2011. Front Microbiol. 2011. PMID: 21847386 Free PMC article.
References
-
- Ahn, J.-H., W.-J. Jang, and G. S. Hayward. 1999. The human cytomegalovirus IE2 and UL112-113 proteins accumulate in viral DNA replication compartments that initiate from the periphery of promyelocytic leukemia protein-associated nuclear bodies (PODs or ND10). J. Virol. 73:10458-10471. - PMC - PubMed
-
- Banks, L., D. Pim, and M. Thomas. 2003. Viruses and the 26S proteasome: hacking into destruction. Trends Biochem. Sci. 28:452-459. - PubMed
-
- Blanchette, P., and P. E. Branton. 2009. Manipulation of the ubiquitin-proteasome pathway by small DNA tumor viruses. Virology 384:317-323. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

