Filling the gap, evolutionarily conserved Omp85 in plastids of chromalveolates
- PMID: 20042599
- PMCID: PMC2825478
- DOI: 10.1074/jbc.M109.074807
Filling the gap, evolutionarily conserved Omp85 in plastids of chromalveolates
Abstract
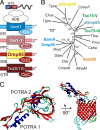
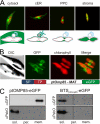
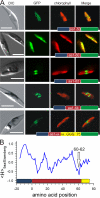
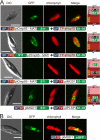
Chromalveolates are a diverse group of protists that include many ecologically and medically relevant organisms such as diatoms and apicomplexan parasites. They possess plastids generally surrounded by four membranes, which evolved by engulfment of a red alga. Today, most plastid proteins must be imported, but many aspects of protein import into complex plastids are still cryptic. In particular, how proteins cross the third outermost membrane has remained unexplained. We identified a protein in the third outermost membrane of the diatom Phaeodactylum tricornutum with properties comparable to those of the Omp85 family. We demonstrate that the targeting route of P. tricornutum Omp85 parallels that of the translocation channel of the outer envelope membrane of chloroplasts, Toc75. In addition, the electrophysiological properties are similar to those of the Omp85 proteins involved in protein translocation. This supports the hypothesis that P. tricornutum Omp85 is involved in precursor protein translocation, which would close a gap in the fundamental understanding of the evolutionary origin and function of protein import in secondary plastids.
Figures

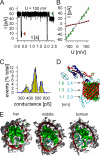
References
-
- Field C. B., Behrenfeld M. J., Randerson J. T., Falkowski P. (1998) Science 281, 237–240 - PubMed
-
- Gould S. B., Waller R. F., McFadden G. I. (2008) Annu. Rev. Plant Biol. 59, 491–517 - PubMed
-
- Bolte K., Bullmann L., Hempel F., Bozarth A., Zauner S., Maier U. G. (2009) J. Eukaryot. Microbiol. 56, 9–15 - PubMed
-
- Hempel F., Bullmann L., Lau J., Zauner S., Maier U. G. (2009) Mol. Biol. Evol. 26, 1781–1790 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

