Energetic frustration of apomyoglobin folding: role of the B helix
- PMID: 20043917
- PMCID: PMC2827033
- DOI: 10.1016/j.jmb.2009.12.040
Energetic frustration of apomyoglobin folding: role of the B helix
Abstract
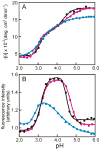
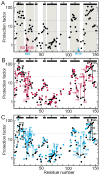
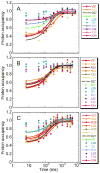
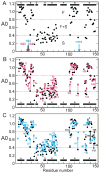
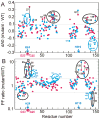
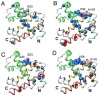
Apomyoglobin folds by a sequential mechanism in which the A, G, and H helix regions undergo rapid collapse to form a compact intermediate onto which the central portion of the B helix subsequently docks. To investigate the factors that frustrate folding, we have made mutations in the N-terminus of the B helix to stabilize helical structure (in the mutant G23A/G25A) and to promote native-like hydrophobic packing interactions with helix G (in the mutant H24L/H119F). The kinetic and equilibrium intermediates of G23A/G25A and H24L/H119F were studied by hydrogen exchange pulse labeling and interrupted hydrogen/deuterium exchange combined with NMR. For both mutants, stabilization of helical structure in the N-terminal region of the B helix is confirmed by increased exchange protection in the equilibrium molten globule states near pH 4. Increased protection is also observed in the GH turn region in the G23A/G25A mutant, suggesting that stabilization of the B helix facilitates native-like interactions with the C-terminal region of helix G. These interactions are further enhanced in H24L/H119F. The kinetic burst phase intermediates of both mutants show increased protection, relative to wild-type protein, of amides in the N-terminus of the B helix and in part of the E helix. Stabilization of the E helix in the intermediate is attributed to direct interactions between E helix residues and the newly stabilized N-terminus of helix B. Stabilization of native packing between the B and G helices in H24L/H119F also favors formation of native-like interactions in the GH turn and between the G and H helices in the ensemble of burst phase intermediates. We conclude that instability at the N-terminus of the B helix of apomyoglobin contributes to the energetic frustration of folding by preventing docking and stabilization of the E helix.
Copyright (c) 2009 Elsevier Ltd. All rights reserved.
Figures
Similar articles
-
Probing the non-native H helix translocation in apomyoglobin folding intermediates.Biochemistry. 2014 Jun 17;53(23):3767-80. doi: 10.1021/bi500478m. Epub 2014 Jun 4. Biochemistry. 2014. PMID: 24857522 Free PMC article.
-
Folding of apomyoglobin: Analysis of transient intermediate structure during refolding using quick hydrogen deuterium exchange and NMR.Proc Jpn Acad Ser B Phys Biol Sci. 2017;93(1):10-27. doi: 10.2183/pjab.93.002. Proc Jpn Acad Ser B Phys Biol Sci. 2017. PMID: 28077807 Free PMC article. Review.
-
Identification of native and non-native structure in kinetic folding intermediates of apomyoglobin.J Mol Biol. 2006 Jan 6;355(1):139-56. doi: 10.1016/j.jmb.2005.10.047. Epub 2005 Nov 8. J Mol Biol. 2006. PMID: 16300787
-
Conformational and dynamic characterization of the molten globule state of an apomyoglobin mutant with an altered folding pathway.Biochemistry. 2001 Dec 4;40(48):14459-67. doi: 10.1021/bi011500n. Biochemistry. 2001. PMID: 11724558
-
The apomyoglobin folding pathway revisited: structural heterogeneity in the kinetic burst phase intermediate.J Mol Biol. 2002 Sep 20;322(3):483-9. doi: 10.1016/s0022-2836(02)00810-0. J Mol Biol. 2002. PMID: 12225742 Review.
Cited by
-
Probing the non-native H helix translocation in apomyoglobin folding intermediates.Biochemistry. 2014 Jun 17;53(23):3767-80. doi: 10.1021/bi500478m. Epub 2014 Jun 4. Biochemistry. 2014. PMID: 24857522 Free PMC article.
-
Folding of apomyoglobin: Analysis of transient intermediate structure during refolding using quick hydrogen deuterium exchange and NMR.Proc Jpn Acad Ser B Phys Biol Sci. 2017;93(1):10-27. doi: 10.2183/pjab.93.002. Proc Jpn Acad Ser B Phys Biol Sci. 2017. PMID: 28077807 Free PMC article. Review.
-
Transient On- and Off-Pathway Protein Folding Intermediate States Characterized with NMR Relaxation Dispersion.J Phys Chem B. 2022 Nov 24;126(46):9539-9548. doi: 10.1021/acs.jpcb.2c05592. Epub 2022 Nov 10. J Phys Chem B. 2022. PMID: 36354189 Free PMC article.
-
Misfolding and amyloid aggregation of apomyoglobin.Int J Mol Sci. 2013 Jul 9;14(7):14287-300. doi: 10.3390/ijms140714287. Int J Mol Sci. 2013. PMID: 23839096 Free PMC article. Review.
-
Photolytic labeling to probe molecular interactions in lyophilized powders.Mol Pharm. 2013 Dec 2;10(12):4629-39. doi: 10.1021/mp4004332. Epub 2013 Oct 29. Mol Pharm. 2013. PMID: 24125175 Free PMC article.
References
-
- Wright PE, Baldwin RL. The folding process of apomyoglobin. In: Pain RH, editor. Mechanisms of Protein Folding. Oxford University Press; Oxford: 2000. pp. 309–329.
-
- Jennings PA, Wright PE. Formation of a molten globule intermediate early in the kinetic folding pathway of apomyoglobin. Science. 1993;262:892–896. - PubMed
-
- Jamin M, Baldwin RL. Two forms of the pH 4 folding intermediate of apomyoglobin. J Mol Biol. 1998;276:491–504. - PubMed
-
- Nishimura C, Dyson HJ, Wright PE. The apomyoglobin folding pathway revisited: structural heterogeneity in the kinetic burst phase intermediate. J Mol Biol. 2002;322:483–489. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

