AraGEM, a genome-scale reconstruction of the primary metabolic network in Arabidopsis
- PMID: 20044452
- PMCID: PMC2815881
- DOI: 10.1104/pp.109.148817
AraGEM, a genome-scale reconstruction of the primary metabolic network in Arabidopsis
Abstract
Genome-scale metabolic network models have been successfully used to describe metabolism in a variety of microbial organisms as well as specific mammalian cell types and organelles. This systems-based framework enables the exploration of global phenotypic effects of gene knockouts, gene insertion, and up-regulation of gene expression. We have developed a genome-scale metabolic network model (AraGEM) covering primary metabolism for a compartmentalized plant cell based on the Arabidopsis (Arabidopsis thaliana) genome. AraGEM is a comprehensive literature-based, genome-scale metabolic reconstruction that accounts for the functions of 1,419 unique open reading frames, 1,748 metabolites, 5,253 gene-enzyme reaction-association entries, and 1,567 unique reactions compartmentalized into the cytoplasm, mitochondrion, plastid, peroxisome, and vacuole. The curation process identified 75 essential reactions with respective enzyme associations not assigned to any particular gene in the Kyoto Encyclopedia of Genes and Genomes or AraCyc. With the addition of these reactions, AraGEM describes a functional primary metabolism of Arabidopsis. The reconstructed network was transformed into an in silico metabolic flux model of plant metabolism and validated through the simulation of plant metabolic functions inferred from the literature. Using efficient resource utilization as the optimality criterion, AraGEM predicted the classical photorespiratory cycle as well as known key differences between redox metabolism in photosynthetic and nonphotosynthetic plant cells. AraGEM is a viable framework for in silico functional analysis and can be used to derive new, nontrivial hypotheses for exploring plant metabolism.
Figures

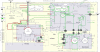


Similar articles
-
AlgaGEM--a genome-scale metabolic reconstruction of algae based on the Chlamydomonas reinhardtii genome.BMC Genomics. 2011 Dec 22;12 Suppl 4(Suppl 4):S5. doi: 10.1186/1471-2164-12-S4-S5. Epub 2011 Dec 22. BMC Genomics. 2011. PMID: 22369158 Free PMC article.
-
Zea mays iRS1563: a comprehensive genome-scale metabolic reconstruction of maize metabolism.PLoS One. 2011;6(7):e21784. doi: 10.1371/journal.pone.0021784. Epub 2011 Jul 6. PLoS One. 2011. PMID: 21755001 Free PMC article.
-
Are we ready for genome-scale modeling in plants?Plant Sci. 2012 Aug;191-192:53-70. doi: 10.1016/j.plantsci.2012.04.010. Epub 2012 May 3. Plant Sci. 2012. PMID: 22682565 Review.
-
C4GEM, a genome-scale metabolic model to study C4 plant metabolism.Plant Physiol. 2010 Dec;154(4):1871-85. doi: 10.1104/pp.110.166488. Epub 2010 Oct 25. Plant Physiol. 2010. PMID: 20974891 Free PMC article.
-
Plant genome-scale metabolic reconstruction and modelling.Curr Opin Biotechnol. 2013 Apr;24(2):271-7. doi: 10.1016/j.copbio.2012.08.007. Epub 2012 Sep 1. Curr Opin Biotechnol. 2013. PMID: 22947602 Review.
Cited by
-
Systematic comparison of C3 and C4 plants based on metabolic network analysis.BMC Syst Biol. 2012;6 Suppl 2(Suppl 2):S9. doi: 10.1186/1752-0509-6-S2-S9. Epub 2012 Dec 12. BMC Syst Biol. 2012. PMID: 23281598 Free PMC article.
-
Identifying genotype-by-environment interactions in the metabolism of germinating arabidopsis seeds using generalized genetical genomics.Plant Physiol. 2013 Jun;162(2):553-66. doi: 10.1104/pp.113.216176. Epub 2013 Apr 19. Plant Physiol. 2013. PMID: 23606598 Free PMC article.
-
Enhancing C3 photosynthesis.Plant Physiol. 2010 Oct;154(2):589-92. doi: 10.1104/pp.110.160952. Plant Physiol. 2010. PMID: 20921190 Free PMC article. No abstract available.
-
Mild reductions in cytosolic NADP-dependent isocitrate dehydrogenase activity result in lower amino acid contents and pigmentation without impacting growth.Amino Acids. 2010 Oct;39(4):1055-66. doi: 10.1007/s00726-010-0617-0. Epub 2010 May 16. Amino Acids. 2010. PMID: 20473773 Free PMC article.
-
Starch biosynthesis in cassava: a genome-based pathway reconstruction and its exploitation in data integration.BMC Syst Biol. 2013 Aug 10;7:75. doi: 10.1186/1752-0509-7-75. BMC Syst Biol. 2013. PMID: 23938102 Free PMC article.
References
-
- Abrol YP, Sawhney SK, Naik MS (1983) Light and dark assimilation of nitrate in plants. Plant Cell Environ 6 595–599
-
- Backhausen JE, Vetter S, Baalmann E, Kitzmann C, Scheibe R (1998) NAD-dependent malate dehydrogenase and glyceraldehyde 3-phosphate dehydrogenase isoenzymes play an important role in dark metabolism of various plastid types. Planta 205 359–366
-
- Bonarius HPJ, Schmid G, Tramper J (1997) Flux analysis of underdetermined metabolic networks: the quest for the missing constraints. Trends Biotechnol 15 308–314
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

