A computational screen for site selective A-to-I editing detects novel sites in neuron specific Hu proteins
- PMID: 20047656
- PMCID: PMC2831006
- DOI: 10.1186/1471-2105-11-6
A computational screen for site selective A-to-I editing detects novel sites in neuron specific Hu proteins
Abstract
Background: Several bioinformatic approaches have previously been used to find novel sites of ADAR mediated A-to-I RNA editing in human. These studies have discovered thousands of genes that are hyper-edited in their non-coding intronic regions, especially in alu retrotransposable elements, but very few substrates that are site-selectively edited in coding regions. Known RNA edited substrates suggest, however, that site selective A-to-I editing is particularly important for normal brain development in mammals.
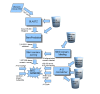
Results: We have compiled a screen that enables the identification of new sites of site-selective editing, primarily in coding sequences. To avoid hyper-edited repeat regions, we applied our screen to the alu-free mouse genome. Focusing on the mouse also facilitated better experimental verification. To identify candidate sites of RNA editing, we first performed an explorative screen based on RNA structure and genomic sequence conservation. We further evaluated the results of the explorative screen by determining which transcripts were enriched for A-G mismatches between the genomic template and the expressed sequence since the editing product, inosine (I), is read as guanosine (G) by the translational machinery. For expressed sequences, we only considered coding regions to focus entirely on re-coding events. Lastly, we refined the results from the explorative screen using a novel scoring scheme based on characteristics for known A-to-I edited sites. The extent of editing in the final candidate genes was verified using total RNA from mouse brain and 454 sequencing.
Conclusions: Using this method, we identified and confirmed efficient editing at one site in the Gabra3 gene. Editing was also verified at several other novel sites within candidates predicted to be edited. Five of these sites are situated in genes coding for the neuron-specific RNA binding proteins HuB and HuD.
Figures


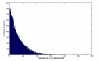
References
-
- Li Q, Lee JA, Black DL. Neuronal regulation of alternative pre-mRNA splicing. Nat Rev Neurosci. 2007;8(11):819–831. - PubMed
-
- Maydanovych O, Beal PA. Breaking the central dogma by RNA editing. Chem Rev. 2006;106(8):3397–3411. - PubMed
-
- Levanon EY, Eisenberg E, Yelin R, Nemzer S, Hallegger M, Shemesh R, Fligelman ZY, Shoshan A, Pollock SR, Sztybel D. Systematic identification of abundant A-to-I editing sites in the human transcriptome. Nat Biotechnol. 2004;22(8):1001–1005. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

