Conserved symbiotic plasmid DNA sequences in the multireplicon pangenomic structure of Rhizobium etli
- PMID: 20048063
- PMCID: PMC2832352
- DOI: 10.1128/AEM.02039-09
Conserved symbiotic plasmid DNA sequences in the multireplicon pangenomic structure of Rhizobium etli
Abstract
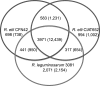
Strains of the same bacterial species often show considerable genomic variation. To examine the extent of such variation in Rhizobium etli, the complete genome sequence of R. etli CIAT652 and the partial genomic sequences of six additional R. etli strains having different geographical origins were determined. The sequences were compared with each other and with the previously reported genome sequence of R. etli CFN42. DNA sequences common to all strains constituted the greater part of these genomes and were localized in both the chromosome and large plasmids. About 700 to 1,000 kb of DNA that did not match sequences of the complete genomes of strains CIAT652 and CFN42 was unique to each R. etli strain. These sequences were distributed throughout the chromosome as individual genes or chromosomal islands and in plasmids, and they encoded accessory functions, such as transport of sugars and amino acids, or secondary metabolism; they also included mobile elements and hypothetical genes. Sequences corresponding to symbiotic plasmids showed high levels of nucleotide identity (about 98 to 99%), whereas chromosomal sequences and the sequences with matches to other plasmids showed lower levels of identity (on average, about 90 to 95%). We concluded that R. etli has a pangenomic structure with a core genome composed of both chromosomal and plasmid sequences, including a highly conserved symbiotic plasmid, despite the overall genomic divergence.
Figures




Similar articles
-
Evolutionary dynamics of insertion sequences in relation to the evolutionary histories of the chromosome and symbiotic plasmid genes of Rhizobium etli populations.Appl Environ Microbiol. 2010 Oct;76(19):6504-13. doi: 10.1128/AEM.01001-10. Epub 2010 Jul 30. Appl Environ Microbiol. 2010. PMID: 20675442 Free PMC article.
-
Diversification of DNA sequences in the symbiotic genome of Rhizobium etli.J Bacteriol. 2005 Nov;187(21):7185-92. doi: 10.1128/JB.187.21.7185-7192.2005. J Bacteriol. 2005. PMID: 16237002 Free PMC article.
-
Housekeeping genes essential for pantothenate biosynthesis are plasmid-encoded in Rhizobium etli and Rhizobium leguminosarum.BMC Microbiol. 2011 Apr 5;11:66. doi: 10.1186/1471-2180-11-66. BMC Microbiol. 2011. PMID: 21463532 Free PMC article.
-
The conjugative plasmid of a bean-nodulating Sinorhizobium fredii strain is assembled from sequences of two Rhizobium plasmids and the chromosome of a Sinorhizobium strain.BMC Microbiol. 2011 Jun 25;11:149. doi: 10.1186/1471-2180-11-149. BMC Microbiol. 2011. PMID: 21702991 Free PMC article.
-
Genomic basis of symbiovar mimosae in Rhizobium etli.BMC Genomics. 2014 Jul 8;15(1):575. doi: 10.1186/1471-2164-15-575. BMC Genomics. 2014. PMID: 25005495 Free PMC article.
Cited by
-
Transfer of the Symbiotic Plasmid of Rhizobium etli CFN42 to Endophytic Bacteria Inside Nodules.Front Microbiol. 2020 Jul 29;11:1752. doi: 10.3389/fmicb.2020.01752. eCollection 2020. Front Microbiol. 2020. PMID: 32849381 Free PMC article.
-
Phylogenomic Rhizobium Species Are Structured by a Continuum of Diversity and Genomic Clusters.Front Microbiol. 2019 Apr 30;10:910. doi: 10.3389/fmicb.2019.00910. eCollection 2019. Front Microbiol. 2019. PMID: 31114559 Free PMC article.
-
Genome sequence of Rhizobium sp. strain CCGE510, a symbiont isolated from nodules of the endangered wild bean Phaseolus albescens.J Bacteriol. 2012 Nov;194(22):6310-1. doi: 10.1128/JB.01536-12. J Bacteriol. 2012. PMID: 23105056 Free PMC article.
-
Genomic studies of nitrogen-fixing rhizobial strains from Phaseolus vulgaris seeds and nodules.BMC Genomics. 2016 Sep 6;17(1):711. doi: 10.1186/s12864-016-3053-z. BMC Genomics. 2016. PMID: 27601031 Free PMC article.
-
Functional conservation of the capacity for ent-kaurene biosynthesis and an associated operon in certain rhizobia.J Bacteriol. 2014 Jan;196(1):100-6. doi: 10.1128/JB.01031-13. Epub 2013 Oct 18. J Bacteriol. 2014. PMID: 24142247 Free PMC article.
References
-
- Abby, S., and V. Daubin. 2007. Comparative genomics and the evolution of prokaryotes. Trends Microbiol. 15:135-141. - PubMed
-
- Andersson, S. G., A. Zomorodipour, J. O. Andersson, T. Sicheritz-Ponten, U. C. Alsmark, R. M. Podowski, A. K. Naslund, A. S. Eriksson, H. H. Winkler, and C. G. Kurland. 1998. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396:133-140. - PubMed
-
- Bentley, S. 2009. Sequencing the species pan-genome. Nat. Rev. Microbiol. 7:258-259. - PubMed
-
- Bentley, S. D., K. F. Chater, A. M. Cerdeno-Tarraga, G. L. Challis, N. R. Thomson, K. D. James, D. E. Harris, M. A. Quail, H. Kieser, D. Harper, A. Bateman, S. Brown, G. Chandra, C. W. Chen, M. Collins, A. Cronin, A. Fraser, A. Goble, J. Hidalgo, T. Hornsby, S. Howarth, C. H. Huang, T. Kieser, L. Larke, L. Murphy, K. Oliver, S. O'Neil, E. Rabbinowitsch, M. A. Rajandream, K. Rutherford, S. Rutter, K. Seeger, D. Saunders, S. Sharp, R. Squares, S. Squares, K. Taylor, T. Warren, A. Wietzorrek, J. Woodward, B. G. Barrell, J. Parkhill, and D. A. Hopwood. 2002. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417:141-147. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

