Commensal microbiota and CD8+ T cells shape the formation of invariant NKT cells
- PMID: 20048124
- PMCID: PMC3458428
- DOI: 10.4049/jimmunol.0902620
Commensal microbiota and CD8+ T cells shape the formation of invariant NKT cells
Abstract
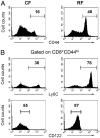
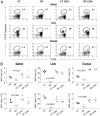
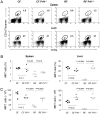
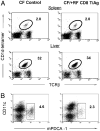
Commensal bacteria play an important role in formation of the immune system, but the mechanisms involved are incompletely understood. In this study, we analyze CD1d-restricted invariant NKT (iNKT) cells in germfree mice and in two colonies of C57BL/6 mice termed conventional flora and restricted flora (RF), stably bearing commensal microbial communities of diverse but distinct composition. In germfree mice, iNKT cells were moderately reduced, suggesting that commensal microbiota were partially required for the antigenic drive in maintaining systemic iNKT cells. Surprisingly, even greater depletion of iNKT cell population occurred in RF mice. This was in part attributable to reduced RF levels of intestinal microbial taxa (Sphingomonas spp.) known to express antigenic glycosphingolipid products. However, memory and activated CD8(+) T cells were also expanded in RF mice, prompting us to test whether CD8(+) T cell activity might be further depleting iNKT cells. Indeed, iNKT cell numbers were restored in RF mice bearing the CD8alpha(-/-) genotype or in adult wild-type RF mice acutely depleted with anti-CD8 Ab. Moreover, iNKT cells were restored in RF mice bearing the Prf1(-/-) phenotype, a key component of cytolytic function. These findings indicate that commensal microbiota, through positive (antigenic drive) and negative (cytolytic depletion by CD8(+) T cells) mechanisms, profoundly shape the iNKT cell compartment. Because individuals greatly vary in the composition of their microbial communities, enteric microbiota may play an important epigenetic role in the striking differences in iNKT cell abundance in humans and therefore in their potential contribution to host immune status.
Figures



References
-
- Savage DC. Microbial ecology of the gastrointestinal tract. Annu Rev Microbiol. 1977;31:107–133. - PubMed
-
- Conway PL. Microbial ecology of the human large intestine. In: Gibson GR, editor. Human Colonic Bacteria: Role in Nutrition, Physiology, And Pathology. CRC Press; Boca Raton, FL: 1995. pp. 1–24.
-
- Mackie RI, Sghir A, Gaskins HR. Developmental microbial ecology of the neonatal gastrointestinal tract. Am J Clin Nutr. 1999;69:1035S–1045S. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- AI52031/AI/NIAID NIH HHS/United States
- GM07185/GM/NIGMS NIH HHS/United States
- CA016042/CA/NCI NIH HHS/United States
- DK69434/DK/NIDDK NIH HHS/United States
- DK349870/DK/NIDDK NIH HHS/United States
- DK46763/DK/NIDDK NIH HHS/United States
- R01 DK069434/DK/NIDDK NIH HHS/United States
- T32 GM007185/GM/NIGMS NIH HHS/United States
- R01 AI078885/AI/NIAID NIH HHS/United States
- AI69296/AI/NIAID NIH HHS/United States
- P01 DK046763/DK/NIDDK NIH HHS/United States
- R01 AI069296/AI/NIAID NIH HHS/United States
- T32 AI052031/AI/NIAID NIH HHS/United States
- P30 CA016042/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

