Structure of D-lactate dehydrogenase from Aquifex aeolicus complexed with NAD(+) and lactic acid (or pyruvate)
- PMID: 20054113
- PMCID: PMC2802865
- DOI: 10.1107/S1744309109044935
Structure of D-lactate dehydrogenase from Aquifex aeolicus complexed with NAD(+) and lactic acid (or pyruvate)
Abstract
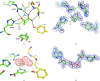
The crystal structure of D-lactate dehydrogenase from Aquifex aeolicus (aq_727) was determined to 2.12 A resolution in space group P2(1)2(1)2(1), with unit-cell parameters a = 90.94, b = 94.43, c = 188.85 A. The structure was solved by molecular replacement using the coenzyme-binding domain of Lactobacillus helveticus D-lactate dehydrogenase and contained two homodimers in the asymmetric unit. Each subunit of the homodimer was found to be in a ;closed' conformation with the NADH cofactor bound to the coenzyme-binding domain and with a lactate (or pyruvate) molecule bound at the interdomain active-site cleft.
Figures


Similar articles
-
Structure of SurE protein from Aquifex aeolicus VF5 at 1.5 A resolution.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009 Dec 1;65(Pt 12):1204-8. doi: 10.1107/S1744309109043814. Epub 2009 Nov 27. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2009. PMID: 20054112 Free PMC article.
-
Domain closure, substrate specificity and catalysis of D-lactate dehydrogenase from Lactobacillus bulgaricus.J Mol Biol. 2002 Apr 19;318(1):109-19. doi: 10.1016/S0022-2836(02)00086-4. J Mol Biol. 2002. PMID: 12054772
-
Crystal structure and thermodynamic properties of d-lactate dehydrogenase from Lactobacillus jensenii.Int J Biol Macromol. 2014 Jul;68:151-7. doi: 10.1016/j.ijbiomac.2014.04.048. Epub 2014 Apr 30. Int J Biol Macromol. 2014. PMID: 24794195
-
Distinct conformation-mediated functions of an active site loop in the catalytic reactions of NAD-dependent D-lactate dehydrogenase and formate dehydrogenase.J Biol Chem. 2005 Apr 29;280(17):17068-75. doi: 10.1074/jbc.M500970200. Epub 2005 Feb 25. J Biol Chem. 2005. PMID: 15734738
-
Engineering d-Lactate Dehydrogenase to Favor an Non-natural Cofactor Nicotinamide Cytosine Dinucleotide.Chembiochem. 2020 Jul 16;21(14):1972-1975. doi: 10.1002/cbic.201900766. Epub 2020 Mar 31. Chembiochem. 2020. PMID: 32175634
Cited by
-
An aldo-keto reductase with 2-keto-l-gulonate reductase activity functions in l-tartaric acid biosynthesis from vitamin C in Vitis vinifera.J Biol Chem. 2019 Nov 1;294(44):15932-15946. doi: 10.1074/jbc.RA119.010196. Epub 2019 Sep 4. J Biol Chem. 2019. PMID: 31488549 Free PMC article.
-
CrystalDock: a novel approach to fragment-based drug design.J Chem Inf Model. 2011 Oct 24;51(10):2573-80. doi: 10.1021/ci200357y. Epub 2011 Oct 5. J Chem Inf Model. 2011. PMID: 21910501 Free PMC article.
-
The D-Lactate Dehydrogenase from Sporolactobacillus inulinus Also Possessing Reversible Deamination Activity.PLoS One. 2015 Sep 23;10(9):e0139066. doi: 10.1371/journal.pone.0139066. eCollection 2015. PLoS One. 2015. PMID: 26398356 Free PMC article.
-
Molecular mechanisms of vancomycin resistance.Protein Sci. 2020 Mar;29(3):654-669. doi: 10.1002/pro.3819. Epub 2020 Jan 23. Protein Sci. 2020. PMID: 31899563 Free PMC article. Review.
-
Classification, substrate specificity and structural features of D-2-hydroxyacid dehydrogenases: 2HADH knowledgebase.BMC Evol Biol. 2018 Dec 22;18(1):199. doi: 10.1186/s12862-018-1309-8. BMC Evol Biol. 2018. PMID: 30577795 Free PMC article.
References
-
- Abola, A., Bernstein, F. C., Bryant, S. H., Koetzle, T. F. & Weng, J. (1987). Crystallographic Databases – Information Content, Software Systems, Scientific Applications, edited by F. H. Allen, G. Bergerhoff & R. Sievers, pp. 107–132. Bonn/Cambridge/Chester: Data Commission of the International Union of Crystallography.
-
- Bradford, M. M. (1976). Anal. Biochem.72, 248–254. - PubMed
-
- DeLano, W. L. (2008). PyMOL Molecular Viewer. DeLano Scientific, Palo Alto, California, USA. http://www.pymol.org.
-
- Ellis, M. J., Antonyuk, S. & Hasnain, S. S. (2002). Acta Cryst. D58, 456–458. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources

