The taming of small heat-shock proteins: crystallization of the alpha-crystallin domain from human Hsp27
- PMID: 20054128
- PMCID: PMC2802880
- DOI: 10.1107/S1744309109044571
The taming of small heat-shock proteins: crystallization of the alpha-crystallin domain from human Hsp27
Abstract
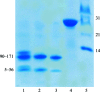
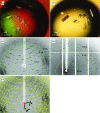
Small heat-shock proteins (sHsps) are ubiquitous molecular chaperones. sHsps function as homooligomers or heterooligomers that are prone to subunit exchange and structural plasticity. Here, a procedure for obtaining diffraction-quality crystals of the alpha-crystallin domain of human Hsp27 is presented. Initially, limited proteolysis was used to delineate the corresponding stable fragment (residues 90-171). This fragment could be crystallized, but examination of the crystals using X-rays indicated partial disorder. The surface-entropy reduction approach was applied to ameliorate the crystal quality. Consequently, a double mutant E125A/E126A of the 90-171 fragment yielded well ordered crystals that diffracted to 2.0 A resolution.
Figures
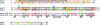
Similar articles
-
Wrapping the alpha-crystallin domain fold in a chaperone assembly.J Mol Biol. 2005 Oct 14;353(1):68-79. doi: 10.1016/j.jmb.2005.08.025. J Mol Biol. 2005. PMID: 16165157
-
Specific sequences in the N-terminal domain of human small heat-shock protein HSPB6 dictate preferential hetero-oligomerization with the orthologue HSPB1.J Biol Chem. 2017 Jun 16;292(24):9944-9957. doi: 10.1074/jbc.M116.773515. Epub 2017 May 9. J Biol Chem. 2017. PMID: 28487364 Free PMC article.
-
Structure and properties of G84R and L99M mutants of human small heat shock protein HspB1 correlating with motor neuropathy.Arch Biochem Biophys. 2013 Oct 1;538(1):16-24. doi: 10.1016/j.abb.2013.07.028. Epub 2013 Aug 12. Arch Biochem Biophys. 2013. PMID: 23948568
-
Small heat shock proteins from extremophiles: a review.Extremophiles. 2004 Feb;8(1):1-11. doi: 10.1007/s00792-003-0362-3. Epub 2003 Nov 19. Extremophiles. 2004. PMID: 15064984 Review.
-
Chaperone-like activity of alpha-crystallin and other small heat shock proteins.Curr Protein Pept Sci. 2001 Sep;2(3):205-25. doi: 10.2174/1389203013381107. Curr Protein Pept Sci. 2001. PMID: 12369933 Review.
Cited by
-
New HSP27 inhibitors efficiently suppress drug resistance development in cancer cells.Oncotarget. 2016 Oct 18;7(42):68156-68169. doi: 10.18632/oncotarget.11905. Oncotarget. 2016. PMID: 27626687 Free PMC article.
-
N- and C-terminal regions of αB-crystallin and Hsp27 mediate inhibition of amyloid nucleation, fibril binding, and fibril disaggregation.J Biol Chem. 2020 Jul 17;295(29):9838-9854. doi: 10.1074/jbc.RA120.012748. Epub 2020 May 16. J Biol Chem. 2020. PMID: 32417755 Free PMC article.
-
Crystal structures of truncated alphaA and alphaB crystallins reveal structural mechanisms of polydispersity important for eye lens function.Protein Sci. 2010 May;19(5):1031-43. doi: 10.1002/pro.380. Protein Sci. 2010. PMID: 20440841 Free PMC article.
References
-
- Arrigo, A. P., Simon, S., Gibert, B., Kretz-Remy, C., Nivon, M., Czekalla, A., Guillet, D., Moulin, M., Diaz-Latoud, C. & Vicart, P. (2007). FEBS Lett.581, 3665–3674. - PubMed
-
- Bagneris, C., Bateman, O. A., Naylor, C. E., Cronin, N., Boelens, W. C., Keep, N. H. & Slingsby, C. (2009). J. Mol. Biol.392, 1242–1252. - PubMed
-
- Bukach, O. V., Seit-Nebi, A. S., Marston, S. B. & Gusev, N. B. (2004). Eur. J. Biochem.271, 291–302. - PubMed
-
- Clark, J. I. & Muchowski, P. J. (2000). Curr. Opin. Struct. Biol.10, 52–59. - PubMed
-
- Diederichs, K. & Karplus, P. A. (1997). Nature Struct. Biol.4, 269–275. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

