Profiling of N-acetylated protein termini provides in-depth insights into the N-terminal nature of the proteome
- PMID: 20061308
- PMCID: PMC2871424
- DOI: 10.1074/mcp.M900463-MCP200
Profiling of N-acetylated protein termini provides in-depth insights into the N-terminal nature of the proteome
Abstract
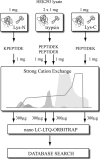
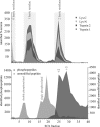
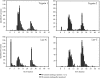
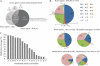
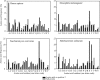
N-terminal processing of proteins is a process affecting a large part of the eukaryotic proteome. Although N-terminal processing is an essential process, not many large inventories are available, in particular not for human proteins. Here we show that by using dedicated mass spectrometry-based proteomics techniques it is possible to unravel N-terminal processing in a semicomprehensive way. Our multiprotease approach led to the identification of 1391 acetylated human protein N termini in HEK293 cells and revealed that the role of the penultimate position on the cleavage efficiency by the methionine aminopeptidases is essentially conserved from Escherichia coli to human. Sequence analysis and comparisons of amino acid frequencies in the data sets of experimentally derived N-acetylated peptides from Drosophila melanogaster, Saccharomyces cerevisiae, and Halobacterium salinarum showed an exceptionally higher frequency of alanine residues at the penultimate position of human proteins, whereas the penultimate position in S. cerevisiae and H. salinarum is predominantly a serine. Genome-wide comparisons revealed that this effect is not related to protein N-terminal processing but can be traced back to characteristics of the genome.
Figures


Similar articles
-
Bioinformatics analysis of a Saccharomyces cerevisiae N-terminal proteome provides evidence of alternative translation initiation and post-translational N-terminal acetylation.J Proteome Res. 2011 Aug 5;10(8):3578-89. doi: 10.1021/pr2002325. Epub 2011 Jun 20. J Proteome Res. 2011. PMID: 21619078
-
Archaeal N-terminal protein maturation commonly involves N-terminal acetylation: a large-scale proteomics survey.J Mol Biol. 2006 Oct 6;362(5):915-24. doi: 10.1016/j.jmb.2006.07.086. Epub 2006 Aug 3. J Mol Biol. 2006. PMID: 16950390
-
Characterization of N-terminal protein modifications in Pseudomonas aeruginosa PA14.J Proteomics. 2015 Jan 30;114:214-25. doi: 10.1016/j.jprot.2014.11.006. Epub 2014 Nov 21. J Proteomics. 2015. PMID: 25464366
-
Protein amino-terminal modifications and proteomic approaches for N-terminal profiling.Curr Opin Chem Biol. 2015 Feb;24:71-9. doi: 10.1016/j.cbpa.2014.10.026. Epub 2014 Nov 15. Curr Opin Chem Biol. 2015. PMID: 25461725 Review.
-
Protein Termini and Their Modifications Revealed by Positional Proteomics.ACS Chem Biol. 2015 Aug 21;10(8):1754-64. doi: 10.1021/acschembio.5b00189. Epub 2015 Jul 6. ACS Chem Biol. 2015. PMID: 26042555 Review.
Cited by
-
The N-end rule pathway and regulation by proteolysis.Protein Sci. 2011 Aug;20(8):1298-345. doi: 10.1002/pro.666. Protein Sci. 2011. PMID: 21633985 Free PMC article. Review.
-
Deletion of cysteine cathepsins B or L yields differential impacts on murine skin proteome and degradome.Mol Cell Proteomics. 2013 Mar;12(3):611-25. doi: 10.1074/mcp.M112.017962. Epub 2012 Dec 10. Mol Cell Proteomics. 2013. PMID: 23233448 Free PMC article.
-
Exploring the accessible conformations of N-terminal acetylated α-synuclein.FEBS Lett. 2013 Apr 17;587(8):1128-38. doi: 10.1016/j.febslet.2013.02.049. Epub 2013 Mar 13. FEBS Lett. 2013. PMID: 23499431 Free PMC article. Review.
-
Perturbation of the yeast N-acetyltransferase NatB induces elevation of protein phosphorylation levels.BMC Genomics. 2010 Dec 2;11:685. doi: 10.1186/1471-2164-11-685. BMC Genomics. 2010. PMID: 21126336 Free PMC article.
-
Quantitation of human metallothionein isoforms: a family of small, highly conserved, cysteine-rich proteins.Mol Cell Proteomics. 2014 Apr;13(4):1020-33. doi: 10.1074/mcp.M113.033373. Epub 2014 Feb 3. Mol Cell Proteomics. 2014. PMID: 24493013 Free PMC article.
References
-
- Bradshaw R. A., Brickey W. W., Walker K. W. (1998) N-terminal processing: the methionine aminopeptidase and N alpha-acetyl transferase families. Trends Biochem. Sci 23, 263–267 - PubMed
-
- Polevoda B., Sherman F. (2000) Nalpha-terminal acetylation of eukaryotic proteins. J. Biol. Chem 275, 36479–36482 - PubMed
-
- Polevoda B., Sherman F. (2003) N-terminal acetyltransferases and sequence requirements for N-terminal acetylation of eukaryotic proteins. J. Mol. Biol 325, 595–622 - PubMed
-
- Frottin F., Martinez A., Peynot P., Mitra S., Holz R. C., Giglione C., Meinnel T. (2006) The proteomics of N-terminal methionine cleavage. Mol. Cell. Proteomics 5, 2336–2349 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

