DNA polymorphisms and haplotype patterns of transcription factors involved in barley endosperm development are associated with key agronomic traits
- PMID: 20064201
- PMCID: PMC2822787
- DOI: 10.1186/1471-2229-10-5
DNA polymorphisms and haplotype patterns of transcription factors involved in barley endosperm development are associated with key agronomic traits
Abstract
Background: Association mapping is receiving considerable attention in plant genetics for its potential to fine map quantitative trait loci (QTL), validate candidate genes, and identify alleles of interest. In the present study association mapping in barley (Hordeum vulgare L.) is investigated by associating DNA polymorphisms with variation in grain quality traits, plant height, and flowering time to gain further understanding of gene functions involved in the control of these traits. We focused on the four loci BLZ1, BLZ2, BPBF and HvGAMYB that play a role in the regulation of B-hordein expression, the major fraction of the barley storage protein. The association was tested in a collection of 224 spring barley accessions using a two-stage mixed model approach.
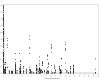
Results: Within the sequenced fragments of four candidate genes we observed different levels of nucleotide diversity. The effect of selection on the candidate genes was tested by Tajima's D which revealed significant values for BLZ1, BLZ2, and BPBF in the subset of two-rowed barleys. Pair-wise LD estimates between the detected SNPs within each candidate gene revealed different intra-genic linkage patterns. On the basis of a more extensive examination of genomic regions surrounding the four candidate genes we found a sharp decrease of LD (r2<0.2 within 1 cM) in all but one flanking regions.Significant marker-trait associations between SNP sites within BLZ1 and flowering time, BPBF and crude protein content and BPBF and starch content were detected. Most haplotypes occurred at frequencies <0.05 and therefore were rejected from the association analysis. Based on haplotype information, BPBF was associated to crude protein content and starch content, BLZ2 showed association to thousand-grain weight and BLZ1 was found to be associated with flowering time and plant height.
Conclusions: Differences in nucleotide diversity and LD pattern within the candidate genes BLZ1, BLZ2, BPBF, and HvGAMYB reflect the impact of selection on the nucleotide sequence of the four candidate loci.Despite significant associations, the analysed candidate genes only explained a minor part of the total genetic variation although they are known to be important factors influencing the expression of seed quality traits. Therefore, we assume that grain quality as well as plant height and flowering time are influenced by many factors each contributing a small part to the expression of the phenotype. A genome-wide association analysis could provide a more comprehensive picture of loci involved in the regulation of grain quality, thousand grain weight and the other agronomic traits that were analyzed in this study. However, despite available high-throughput genotyping arrays the marker density along the barely genome is still insufficient to cover all associations in a whole genome scan. Therefore, the candidate gene-based approach will further play an important role in barley association studies.
Figures
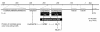

References
-
- Pflieger S, Lefebvre V, Causse M. The candidate gene approach in plant genetics: a review. Mol Breed. 2001;7:275–291. doi: 10.1023/A:1011605013259. - DOI
-
- Bartels D, Thompson RD. Synthesis of messenger-RNAs coding for abundant endosperm proteins during wheat-grain development. Plant Sci. 1986;46:117–125. doi: 10.1016/0168-9452(86)90118-4. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

