Regulation of expression of stromal-derived factor-1 receptors: CXCR4 and CXCR7 in human rhabdomyosarcomas
- PMID: 20068066
- PMCID: PMC2943743
- DOI: 10.1158/1541-7786.MCR-09-0259
Regulation of expression of stromal-derived factor-1 receptors: CXCR4 and CXCR7 in human rhabdomyosarcomas
Abstract
Rhabdomyosarcomas (RMS) express CXCR4 and CXCR7 receptors that bind prometastatic alpha-chemokine stromal-derived factor-1 (SDF-1). In this report, we analyzed the activity of both promoters in a model of less metastatic human embryonal-RMS cell line (RD) and more metastatic alveolar-like RMS (RD cells transduced with paired box gene 3/forkhead homologue; PAX3-FKHR fusion gene). First, CXCR4 is barely detectable in RD and becomes upregulated in RD/PAX3-FKHR cells. In contrast, CXCR7 highly expressed in RD becomes downregulated in RD/PAX3-FKHR cells. Next, promoter deletion and mutation studies revealed that whereas (a) expression of CXCR4 in RD and RD/PAX3-FKHR cells required nuclear respiratory factor-1 (NRF-1) binding site and (b) was additionally upregulated by direct interaction of NRF-1 with PAX3-FKHR, CXCR7 promoter activity required a proximal nuclear factor-kappaB-binding motif. The requirement of these factors for CXCR4 and CXCR7 promoter activities was additionally supported after blocking NRF-1 and nuclear factor-kappaB. Furthermore, CXCR4 expression in PAX3-FKHR(+) RMS cells seems to be enhanced because of the interaction of PAX3-FKHR and NRF-1 proteins in the proximal part of the promoter that prevents access of the negative regulator of transcription YY1 to its binding site. Finally, although hypoxia enhances CXCR4 and CXCR7 promoter activity and receptor expression in RD cells, it inhibits CXCR7 expression in RD/PAX3-FKHR cells. In conclusion, SDF-1 binding receptors CXCR4 and CXCR7 are differently regulated in RMS cells. The upregulation of CXCR4 and downregulation of CXCR7 expression by PAX3-FKHR or hypoxia may give SDF-1 an advantage to better engage the CXCR4 receptor, thus increasing RMS motility.
Figures


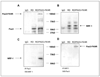



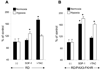
References
-
- Barr FG, Galili N, Holick J, Biegel JA, Rovera G, Emanuel BS. Rearrangement of the PAX3 paired box gene in the paediatric solid tumour alveolar rhabdomyosarcoma. Nat Genet. 1993;3:113–117. - PubMed
-
- Davis RJ, D'Cruz CM, Lovell MA, Biegel JA, Barr FG. Fusion of PAX7 to FKHR by the variant t(1;13)(p36;q14) translocation in alveolar rhabdomyosarcoma. Cancer Res. 1994;54:2869–2872. - PubMed
-
- Hazelton BJ, Houghton JA, Parham DM, et al. Characterization of cell lines derived from xenografts of childhood rhabdomyosarcoma. Cancer Res. 1987;47:4501–4507. - PubMed
-
- Ruymann FB, Newton WA, Jr, Ragab AH, Donaldson MH, Foulkes M. Bone marrow metastases at diagnosis in children and adolescents with rhabdomyosarcoma. A report from the intergroup rhabdomyosarcoma study. Cancer. 1984;53:368–373. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

