Molecular simulation of ab initio protein folding for a millisecond folder NTL9(1-39)
- PMID: 20070076
- PMCID: PMC2835335
- DOI: 10.1021/ja9090353
Molecular simulation of ab initio protein folding for a millisecond folder NTL9(1-39)
Abstract
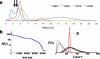
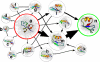
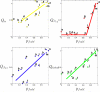
To date, the slowest-folding proteins folded ab initio by all-atom molecular dynamics simulations have had folding times in the range of nanoseconds to microseconds. We report simulations of several folding trajectories of NTL9(1-39), a protein which has a folding time of approximately 1.5 ms. Distributed molecular dynamics simulations in implicit solvent on GPU processors were used to generate ensembles of trajectories out to approximately 40 micros for several temperatures and starting states. At a temperature less than the melting point of the force field, we observe a small number of productive folding events, consistent with predictions from a model of parallel uncoupled two-state simulations. The posterior distribution of the folding rate predicted from the data agrees well with the experimental folding rate (approximately 640/s). Markov State Models (MSMs) built from the data show a gap in the implied time scales indicative of two-state folding and heterogeneous pathways connecting diffuse mesoscopic substates. Structural analysis of the 14 out of 2000 macrostates transited by the top 10 folding pathways reveals that native-like pairing between strands 1 and 2 only occurs for macrostates with p(fold) > 0.5, suggesting beta(12) hairpin formation may be rate-limiting. We believe that using simulation data such as these to seed adaptive resampling simulations will be a promising new method for achieving statistically converged descriptions of folding landscapes at longer time scales than ever before.
Figures
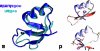
Similar articles
-
Folding processes of the B domain of protein A to the native state observed in all-atom ab initio folding simulations.J Chem Phys. 2008 Jun 21;128(23):235105. doi: 10.1063/1.2937135. J Chem Phys. 2008. PMID: 18570534 Free PMC article.
-
Molecular dynamics with the united-residue model of polypeptide chains. II. Langevin and Berendsen-bath dynamics and tests on model alpha-helical systems.J Phys Chem B. 2005 Jul 21;109(28):13798-810. doi: 10.1021/jp058007w. J Phys Chem B. 2005. PMID: 16852728 Free PMC article.
-
Improved Estimates of Folding Stabilities and Kinetics with Multiensemble Markov Models.Biochemistry. 2024 Nov 19;63(22):3045-3056. doi: 10.1021/acs.biochem.4c00573. Epub 2024 Nov 7. Biochemistry. 2024. PMID: 39509176
-
Simulating the peptide folding kinetic related spectra based on the Markov State Model.Adv Exp Med Biol. 2014;805:199-220. doi: 10.1007/978-3-319-02970-2_9. Adv Exp Med Biol. 2014. PMID: 24446363 Review.
-
Markov state models provide insights into dynamic modulation of protein function.Acc Chem Res. 2015 Feb 17;48(2):414-22. doi: 10.1021/ar5002999. Epub 2015 Jan 3. Acc Chem Res. 2015. PMID: 25625937 Free PMC article. Review.
Cited by
-
Emergence of glass-like behavior in Markov state models of protein folding dynamics.J Am Chem Soc. 2013 Apr 17;135(15):5501-4. doi: 10.1021/ja4002663. Epub 2013 Apr 3. J Am Chem Soc. 2013. PMID: 23540906 Free PMC article.
-
BAD-NEUS: Rapidly converging trajectory stratification.J Chem Phys. 2024 Aug 28;161(8):084109. doi: 10.1063/5.0215975. J Chem Phys. 2024. PMID: 39185846 Free PMC article.
-
Protein folding is mechanistically robust.Biophys J. 2012 Feb 22;102(4):859-67. doi: 10.1016/j.bpj.2012.01.028. Epub 2012 Feb 21. Biophys J. 2012. PMID: 22385857 Free PMC article.
-
Free Energy Landscape and Conformational Kinetics of Hoogsteen Base Pairing in DNA vs. RNA.Biophys J. 2020 Oct 20;119(8):1568-1579. doi: 10.1016/j.bpj.2020.08.031. Epub 2020 Sep 2. Biophys J. 2020. PMID: 32946766 Free PMC article.
-
Dissecting the Energetics of Intrinsically Disordered Proteins via a Hybrid Experimental and Computational Approach.J Phys Chem B. 2019 Dec 12;123(49):10394-10402. doi: 10.1021/acs.jpcb.9b08323. Epub 2019 Dec 3. J Phys Chem B. 2019. PMID: 31702919 Free PMC article.
References
-
- Dill KA, Ozkan SB, Weikl TR, Chodera JD, Voelz VA. Current Opinion in Structural Biology. 2007;17:342–346. - PubMed
-
- Plaxco KW, Simons KT, Baker D. Journal of Molecular Biology. 1998;277:985–994. - PubMed
-
- Yang WY, Gruebele M. Nature. 2003;423:193–197. - PubMed
-
- Kubelka J, Chiu TK, Davies DR, Eaton WA, Hofrichter J. Journal of Molecular Biology. 2006;359:546–553. - PubMed
-
- Kubelka J, Hofrichter J, Eaton WA. Current Opinion in Structural Biology. 2004;14:76–88. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

