Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate
- PMID: 20072126
- PMCID: PMC2884187
- DOI: 10.1038/nature08750
Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate
Abstract
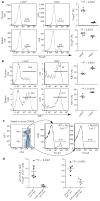
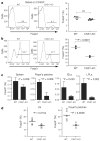
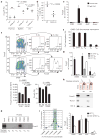
Immune homeostasis is dependent on tight control over the size of a population of regulatory T (T(reg)) cells capable of suppressing over-exuberant immune responses. The T(reg) cell subset is comprised of cells that commit to the T(reg) lineage by upregulating the transcription factor Foxp3 either in the thymus (tT(reg)) or in the periphery (iT(reg)). Considering a central role for Foxp3 in T(reg) cell differentiation and function, we proposed that conserved non-coding DNA sequence (CNS) elements at the Foxp3 locus encode information defining the size, composition and stability of the T(reg) cell population. Here we describe the function of three Foxp3 CNS elements (CNS1-3) in T(reg) cell fate determination in mice. The pioneer element CNS3, which acts to potently increase the frequency of T(reg) cells generated in the thymus and the periphery, binds c-Rel in in vitro assays. In contrast, CNS1, which contains a TGF-beta-NFAT response element, is superfluous for tT(reg) cell differentiation, but has a prominent role in iT(reg) cell generation in gut-associated lymphoid tissues. CNS2, although dispensable for Foxp3 induction, is required for Foxp3 expression in the progeny of dividing T(reg) cells. Foxp3 binds to CNS2 in a Cbf-beta-Runx1 and CpG DNA demethylation-dependent manner, suggesting that Foxp3 recruitment to this 'cellular memory module' facilitates the heritable maintenance of the active state of the Foxp3 locus and, therefore, T(reg) lineage stability. Together, our studies demonstrate that the composition, size and maintenance of the T(reg) cell population are controlled by Foxp3 CNS elements engaged in response to distinct cell-extrinsic or -intrinsic cues.
Conflict of interest statement
Figures

Similar articles
-
c-Rel: a pioneer in directing regulatory T-cell lineage commitment?Eur J Immunol. 2010 Mar;40(3):664-7. doi: 10.1002/eji.201040372. Eur J Immunol. 2010. PMID: 20162555 Review.
-
To be or not to be a Treg cell: lineage decisions controlled by epigenetic mechanisms.Sci Signal. 2011 Feb 1;4(158):pe4. doi: 10.1126/scisignal.2001783. Sci Signal. 2011. PMID: 21285410
-
Smad3 binding to the foxp3 enhancer is dispensable for the development of regulatory T cells with the exception of the gut.J Exp Med. 2012 Aug 27;209(9):1529-35. doi: 10.1084/jem.20112646. Epub 2012 Aug 20. J Exp Med. 2012. PMID: 22908322 Free PMC article.
-
Foxp3 enhancers synergize to maximize regulatory T cell suppressive capacity.J Exp Med. 2021 Aug 2;218(8):e20202415. doi: 10.1084/jem.20202415. Epub 2021 Jun 4. J Exp Med. 2021. PMID: 34086055 Free PMC article.
-
Human regulatory T cells: a unique, stable thymic subset or a reversible peripheral state of differentiation?Immunol Lett. 2007 Nov 30;114(1):9-15. doi: 10.1016/j.imlet.2007.08.012. Epub 2007 Sep 29. Immunol Lett. 2007. PMID: 17945352 Free PMC article. Review.
Cited by
-
Post-Translational Regulations of Foxp3 in Treg Cells and Their Therapeutic Applications.Front Immunol. 2021 Apr 12;12:626172. doi: 10.3389/fimmu.2021.626172. eCollection 2021. Front Immunol. 2021. PMID: 33912156 Free PMC article. Review.
-
Context and location dependence of adaptive Foxp3(+) regulatory T cell formation during immunopathological conditions.Cell Immunol. 2012 Sep;279(1):60-5. doi: 10.1016/j.cellimm.2012.09.009. Epub 2012 Oct 1. Cell Immunol. 2012. PMID: 23089195 Free PMC article. Review.
-
TGFβ control of immune responses in cancer: a holistic immuno-oncology perspective.Nat Rev Immunol. 2023 Jun;23(6):346-362. doi: 10.1038/s41577-022-00796-z. Epub 2022 Nov 15. Nat Rev Immunol. 2023. PMID: 36380023 Free PMC article. Review.
-
ICOS+ Tregs: A Functional Subset of Tregs in Immune Diseases.Front Immunol. 2020 Aug 28;11:2104. doi: 10.3389/fimmu.2020.02104. eCollection 2020. Front Immunol. 2020. PMID: 32983168 Free PMC article. Review.
-
Cutting edge: epigenetic regulation of Foxp3 defines a stable population of CD4+ regulatory T cells in tumors from mice and humans.J Immunol. 2015 Feb 1;194(3):878-82. doi: 10.4049/jimmunol.1402725. Epub 2014 Dec 29. J Immunol. 2015. PMID: 25548231 Free PMC article.
References
-
- Sakaguchi S, Yamaguchi T, Nomura T, Ono M. Regulatory T cells and immune tolerance. Cell. 2008;133:775–787. - PubMed
-
- Zheng Y, Rudensky AY. Foxp3 in control of the regulatory T cell lineage. Nature Immunol. 2007;8:457–462. - PubMed
-
- Gavin MA, et al. Foxp3-dependent programme of regulatory T-cell differentiation. Nature. 2007;445:771–775. - PubMed
-
- Williams LM, Rudensky AY. Maintenance of the Foxp3-dependent developmental program in mature regulatory T cells requires continued expression of Foxp3. Nature Immunol. 2007;8:277–284. - PubMed
-
- Ruthenburg AJ, Allis CD, Wysocka J. Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol Cell. 2007;25:15–30. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

