HJURP binds CENP-A via a highly conserved N-terminal domain and mediates its deposition at centromeres
- PMID: 20080577
- PMCID: PMC2824361
- DOI: 10.1073/pnas.0913709107
HJURP binds CENP-A via a highly conserved N-terminal domain and mediates its deposition at centromeres
Abstract
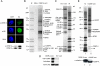
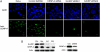
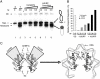
The human histone H3 variant, CENP-A, replaces the conventional histone H3 in centromeric chromatin and, together with centromere-specific DNA-binding factors, directs the assembly of the kinetochore. We purified the prenucelosomal e-CENP-A complex. We found that HJURP, a member of the complex, was required for cell cycle specific targeting of CENP-A to centromeres. HJURP facilitated efficient deposition of CENP-A/H4 tetramers to naked DNA in vitro. Bacterially expressed HJURP binds at a stoichiometric ratio to the CENP-A/H4 tetramer but not to the H3/H4 tetramer. The binding occurred through a conserved HJURP short N-terminal domain, termed CBD. The novel characteristic identified in vertebrates that we named TLTY box of CBD, was essential for formation of the HJURP-CENP-A/H4 complex. Our data identified HJURP as a vertebrate CENP-A chaperone and dissected its mode of interactions with CENP-A.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Mitelman F. Catalog of Chromosome Aberrations in Cancer. 5th Ed. New York: Wiley; 1994.
-
- Henikoff S, Dalal Y. Centromeric chromatin: What makes it unique? Curr Opin Genet Dev. 2005;15:177–184. - PubMed
-
- Cooper JL, Henikoff S. Adaptive evolution of the histone fold domain in centromeric histones. Mol Biol Evol. 2004;21:1712–1718. - PubMed
-
- Smith MM. Centromeres and variant histones: What, where, when, and why? Curr Opin Cell Biol. 2002;14:279–285. - PubMed
-
- Earnshaw WC, Migeon BR. Three related centromere proteins are absent from the inactive centromere of a stable isodicentric chromosome. Chromosoma. 1985;92:290–296. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

