Three conformational snapshots of the hepatitis C virus NS3 helicase reveal a ratchet translocation mechanism
- PMID: 20080715
- PMCID: PMC2818896
- DOI: 10.1073/pnas.0913380107
Three conformational snapshots of the hepatitis C virus NS3 helicase reveal a ratchet translocation mechanism
Abstract
A virally encoded superfamily-2 (SF2) helicase (NS3h) is essential for the replication of hepatitis C virus, a leading cause of liver disease worldwide. Efforts to elucidate the function of NS3h and to develop inhibitors against it, however, have been hampered by limited understanding of its molecular mechanism. Here we show x-ray crystal structures for a set of NS3h complexes, including ground-state and transition-state ternary complexes captured with ATP mimics (ADP.BeF(3) and ). These structures provide, for the first time, three conformational snapshots demonstrating the molecular basis of action for a SF2 helicase. Upon nucleotide binding, overall domain rotation along with structural transitions in motif V and the bound DNA leads to the release of one base from the substrate base-stacking row and the loss of several interactions between NS3h and the 3' DNA segment. As nucleotide hydrolysis proceeds into the transition state, stretching of a "spring" helix and another overall conformational change couples rearrangement of the (d)NTPase active site to additional hydrogen-bonding between NS3h and DNA. Together with biochemistry, these results demonstrate a "ratchet" mechanism involved in the unidirectional translocation and define the step size of NS3h as one base per nucleotide hydrolysis cycle. These findings suggest feasible strategies for developing specific inhibitors to block the action of this attractive, yet largely unexplored drug target.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
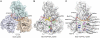
 and ssDNA (dT6). The structures are represented by ribbons and transparent surfaces. The DNA bases and deoxyribose groups are shown with sticks and numerically labeled. The DNA phosphodiester backbones are simplified as orange tubes. The DNA atoms are color coded according to elements. The helicase motifs are color coded in the surface and ribbon respectively in (B) and (C). The distances between the Cα atoms of K244 (domain 1) and S403 (domain 2) are noted. ADP·BeF3 and
and ssDNA (dT6). The structures are represented by ribbons and transparent surfaces. The DNA bases and deoxyribose groups are shown with sticks and numerically labeled. The DNA phosphodiester backbones are simplified as orange tubes. The DNA atoms are color coded according to elements. The helicase motifs are color coded in the surface and ribbon respectively in (B) and (C). The distances between the Cα atoms of K244 (domain 1) and S403 (domain 2) are noted. ADP·BeF3 and  are shown with sticks and color coded.
are shown with sticks and color coded.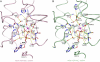
 . NS3h-ADP·BeF3-ssDNA(dT6) is pink and
. NS3h-ADP·BeF3-ssDNA(dT6) is pink and  is green. The non-carbon atoms are color coded according to elements. Protein main chains are simplified as tubes. The main-chain atoms and side chains involved in the coordination of nucleotides are represented by sticks. These protein residues are labeled with single-letter codes, residue numbers and motif codes. The metal ions (magenta) and water molecules (red) are spheres. The interactions are represented by dashed lines.
is green. The non-carbon atoms are color coded according to elements. Protein main chains are simplified as tubes. The main-chain atoms and side chains involved in the coordination of nucleotides are represented by sticks. These protein residues are labeled with single-letter codes, residue numbers and motif codes. The metal ions (magenta) and water molecules (red) are spheres. The interactions are represented by dashed lines.
 (green) complexes, aligned through their ADP moieties. (D) The spring helices from the ground-state and transition-state complexes are aligned through motif Y and the adjacent residues colored in white. Y241 is shown in sticks. (E) Structural remodeling of the (d)NTPase active site from the ground state to the transition state. The complexes are aligned as those in (B). Motif II is highlighted with yellow spheres (carbon atoms).
(green) complexes, aligned through their ADP moieties. (D) The spring helices from the ground-state and transition-state complexes are aligned through motif Y and the adjacent residues colored in white. Y241 is shown in sticks. (E) Structural remodeling of the (d)NTPase active site from the ground state to the transition state. The complexes are aligned as those in (B). Motif II is highlighted with yellow spheres (carbon atoms).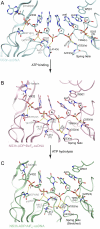
 complex. The carbon elements of the complexes are color coded as in Fig 3. Protein main chains are simplified as tubes and ribbons. The residues involved in ssDNA binding are shown with sticks and color coded according to elements. Side chains are omitted for those residues that interact with ssDNA only through their main-chain atoms. The DNA strands are modeled with sticks and color coded according to elements. The orientations of the DNA bases are noted as either anti or syn. The deoxyribose groups labeled B are in a C2′-endo, whereas the ones labeled A are in C3′-endo. The yellow dashed lines represent hydrogen bonds. The location of the spring helix is noted.
complex. The carbon elements of the complexes are color coded as in Fig 3. Protein main chains are simplified as tubes and ribbons. The residues involved in ssDNA binding are shown with sticks and color coded according to elements. Side chains are omitted for those residues that interact with ssDNA only through their main-chain atoms. The DNA strands are modeled with sticks and color coded according to elements. The orientations of the DNA bases are noted as either anti or syn. The deoxyribose groups labeled B are in a C2′-endo, whereas the ones labeled A are in C3′-endo. The yellow dashed lines represent hydrogen bonds. The location of the spring helix is noted.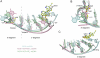
 complexes. NS3h-ssDNA(dA6) and NS3h-ADP·BeF3-ssDNA(dT12) are aligned through domain 2. NS3h-ADP·BeF3-ssDNA(dT12) and
complexes. NS3h-ssDNA(dA6) and NS3h-ADP·BeF3-ssDNA(dT12) are aligned through domain 2. NS3h-ADP·BeF3-ssDNA(dT12) and  are aligned through the ADP moieties. (B) A closer view of the 5′ segments of ssDNA and the β-hairpin. The structures are aligned through domain 2. (C) Structural comparison of the 3′ segments of ssDNA. The structures are aligned through phosphate 1 to 3 of the DNA strands. The complexes are color coded as in Fig 3. The DNA bases and deoxyribose groups are represented by sticks and numerically labeled in each panel. The phosphodiester backbones are simplified as tubes. Protein side chains are modeled with sticks. The carbon atoms in protein residues are highlighted by yellow spheres.
are aligned through the ADP moieties. (B) A closer view of the 5′ segments of ssDNA and the β-hairpin. The structures are aligned through domain 2. (C) Structural comparison of the 3′ segments of ssDNA. The structures are aligned through phosphate 1 to 3 of the DNA strands. The complexes are color coded as in Fig 3. The DNA bases and deoxyribose groups are represented by sticks and numerically labeled in each panel. The phosphodiester backbones are simplified as tubes. Protein side chains are modeled with sticks. The carbon atoms in protein residues are highlighted by yellow spheres.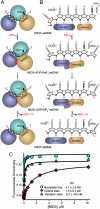
Similar articles
-
Investigation of translocation, DNA unwinding, and protein displacement by NS3h, the helicase domain from the hepatitis C virus helicase.Biochemistry. 2010 Mar 16;49(10):2097-109. doi: 10.1021/bi901977k. Biochemistry. 2010. PMID: 20108974 Free PMC article.
-
The Spring α-Helix Coordinates Multiple Modes of HCV (Hepatitis C Virus) NS3 Helicase Action.J Biol Chem. 2016 Jul 8;291(28):14499-509. doi: 10.1074/jbc.M115.704379. Epub 2016 May 12. J Biol Chem. 2016. PMID: 27226535 Free PMC article.
-
The macroscopic rate of nucleic acid translocation by hepatitis C virus helicase NS3h is dependent on both sugar and base moieties.J Mol Biol. 2010 Jul 16;400(3):354-78. doi: 10.1016/j.jmb.2010.04.065. Epub 2010 May 6. J Mol Biol. 2010. PMID: 20451531 Free PMC article.
-
Step-by-step progress toward understanding the hepatitis C virus RNA helicase.Hepatology. 2006 Jun;43(6):1392-5. doi: 10.1002/hep.21200. Hepatology. 2006. PMID: 16729317 Free PMC article. Review.
-
Structure and function of hepatitis C virus NS3 helicase.Curr Top Microbiol Immunol. 2000;242:171-96. doi: 10.1007/978-3-642-59605-6_9. Curr Top Microbiol Immunol. 2000. PMID: 10592661 Review.
Cited by
-
Common Mechanism of Activated Catalysis in P-loop Fold Nucleoside Triphosphatases-United in Diversity.Biomolecules. 2022 Sep 22;12(10):1346. doi: 10.3390/biom12101346. Biomolecules. 2022. PMID: 36291556 Free PMC article.
-
Comparative structural analysis of human DEAD-box RNA helicases.PLoS One. 2010 Sep 30;5(9):e12791. doi: 10.1371/journal.pone.0012791. PLoS One. 2010. PMID: 20941364 Free PMC article.
-
Impact of bound ssRNA length on allostery in the Dengue Virus NS3 helicase.Nucleic Acids Res. 2023 Nov 10;51(20):11213-11224. doi: 10.1093/nar/gkad808. Nucleic Acids Res. 2023. PMID: 37823592 Free PMC article.
-
Hepatitis C virus RNA replication and assembly: living on the fat of the land.Cell Host Microbe. 2014 Nov 12;16(5):569-79. doi: 10.1016/j.chom.2014.10.008. Epub 2014 Nov 12. Cell Host Microbe. 2014. PMID: 25525790 Free PMC article. Review.
-
Single-molecule imaging reveals the translocation and DNA looping dynamics of hepatitis C virus NS3 helicase.Protein Sci. 2017 Jul;26(7):1391-1403. doi: 10.1002/pro.3136. Epub 2017 Mar 6. Protein Sci. 2017. PMID: 28176403 Free PMC article.
References
-
- De Francesco R, Migliaccio G. Challenges and successes in developing new therapies for hepatitis C. Nature. 2005;436:953–960. - PubMed
-
- Moradpour D, Penin F, Rice CM. Replication of hepatitis C virus. Nat Rev Microbiol. 2007;5:453–463. - PubMed
-
- Kim DW, Gwack Y, Han JH, Choe J. C-terminal domain of the hepatitis C virus NS3 protein contains an RNA helicase activity. Biochem Biophys Res Commun. 1995;215:160–166. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

