Amot recognizes a juxtanuclear endocytic recycling compartment via a novel lipid binding domain
- PMID: 20080965
- PMCID: PMC2852970
- DOI: 10.1074/jbc.M109.096230
Amot recognizes a juxtanuclear endocytic recycling compartment via a novel lipid binding domain
Abstract
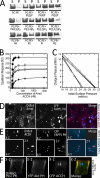
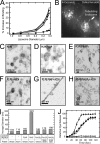
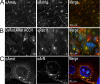
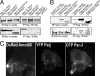
Polarity proteins promote the asymmetric organization of cells by orienting intracellular sorting mechanisms, such as protein trafficking and cytoskeletal assembly. The localization of individual polarity proteins in turn is often determined by association with factors that mediate contact with other cells or the substratum. This arrangement for the Par and Crb apical polarity complexes at the tight junction is disrupted by the adaptor protein Amot. Amot directly binds the scaffolding proteins Patj and Mupp1 and redistributes them and their binding partners from the plasma membrane to endosomes. However, the mechanism by which Amot is targeted to endosomes is unknown. Here, a novel lipid binding domain within Amot is shown to selectively bind with high affinity to membranes containing monophosphorylated phosphatidylinositols and cholesterol. With similar lipid specificity, Amot inserts into and tubulates membranes in vitro and enlarges perinuclear endosomal compartments in cells. Based on the similar distribution of Amot with cholesterol, Rab11, and Arf6, such membrane interactions are identified at juxtanuclear endocytic recycling compartments. Taken together, these findings indicate that Amot is targeted along with associated apical polarity proteins to the endocytic recycling compartment via this novel membrane binding domain.
Figures


Similar articles
-
A Rich1/Amot complex regulates the Cdc42 GTPase and apical-polarity proteins in epithelial cells.Cell. 2006 May 5;125(3):535-48. doi: 10.1016/j.cell.2006.02.045. Cell. 2006. PMID: 16678097
-
Molecular characterization of angiomotin/JEAP family proteins: interaction with MUPP1/Patj and their endogenous properties.Genes Cells. 2007 Apr;12(4):473-86. doi: 10.1111/j.1365-2443.2007.01066.x. Genes Cells. 2007. PMID: 17397395
-
Sorting of membrane and fluid at the apical pole of polarized Madin-Darby canine kidney cells.Mol Biol Cell. 2000 Jun;11(6):2131-50. doi: 10.1091/mbc.11.6.2131. Mol Biol Cell. 2000. PMID: 10848634 Free PMC article.
-
Polarized endocytic transport: the roles of Rab11 and Rab11-FIPs in regulating cell polarity.Histol Histopathol. 2009 Sep;24(9):1171-80. doi: 10.14670/HH-24.1171. Histol Histopathol. 2009. PMID: 19609864 Free PMC article. Review.
-
Lipids in endocytic membrane transport and sorting.Curr Opin Cell Biol. 2003 Aug;15(4):382-8. doi: 10.1016/s0955-0674(03)00078-4. Curr Opin Cell Biol. 2003. PMID: 12892777 Review.
Cited by
-
EPEC effector EspF promotes Crumbs3 endocytosis and disrupts epithelial cell polarity.Cell Microbiol. 2017 Nov;19(11):10.1111/cmi.12757. doi: 10.1111/cmi.12757. Epub 2017 Jul 27. Cell Microbiol. 2017. PMID: 28618099 Free PMC article.
-
Dynamic Polarization of Rab11a Modulates Crb2a Localization and Impacts Signaling to Regulate Retinal Neurogenesis.Front Cell Dev Biol. 2021 Feb 9;8:608112. doi: 10.3389/fcell.2020.608112. eCollection 2020. Front Cell Dev Biol. 2021. PMID: 33634099 Free PMC article.
-
Compensatory Role of Inositol 5-Phosphatase INPP5B to OCRL in Primary Cilia Formation in Oculocerebrorenal Syndrome of Lowe.PLoS One. 2013 Jun 21;8(6):e66727. doi: 10.1371/journal.pone.0066727. Print 2013. PLoS One. 2013. PMID: 23805271 Free PMC article.
-
Determining the minimal amount of DMSO necessary to stabilize the Angiomotin lipid binding domain.Indiana Univ J Undergrad Res. 2024;8:10.14434/iujur.v8i1.31200. doi: 10.14434/iujur.v8i1.31200. Epub 2024 Jun 13. Indiana Univ J Undergrad Res. 2024. PMID: 39866826 Free PMC article.
-
The Angiomotins--from discovery to function.FEBS Lett. 2014 Aug 19;588(16):2693-703. doi: 10.1016/j.febslet.2014.02.006. Epub 2014 Feb 15. FEBS Lett. 2014. PMID: 24548561 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

