Reversibility of beta-amyloid self-assembly: effects of pH and added salts assessed by fluorescence photobleaching recovery
- PMID: 20085314
- PMCID: PMC2859319
- DOI: 10.1021/bm900833b
Reversibility of beta-amyloid self-assembly: effects of pH and added salts assessed by fluorescence photobleaching recovery
Abstract
The 40-residue peptide isoform beta-amyloid (Abeta(1-40)) is associated with Alzheimer's disease. Although found in the tangles and fibrous mats that characterize the brain in advanced stages of the disease, the toxic form of Abeta is believed to be oligomers or "protofibrils". Characterization of these fairly small structures in solution, especially in the presence of the much larger assemblies they also form, is a daunting task. Additionally, little is known about the rate of Abeta assembly or whether it can be triggered easily. Perhaps most importantly, the conditions for reversing assembly are not fully understood. Fluorescence photobleaching with modulation detection of the recovery profile is a sensitive and materials-efficient way to measure diffusers over a wide range of hydrodynamic sizes. The method does require attachment of a fluorescent label. Experiments to validate the use of 5-carboxyfluorescein-labeled Abeta(1-40) as a representative of the unlabeled, naturally occurring material included variation of photobleaching time and mixture of labeled and unlabeled materials. A dialysis cell facilitated rapid in situ changes in pH and salt conditions. Multiple steps and complex protocols can be explored with relative ease. Oligomeric aggregates were found by fluorescence photobleaching recovery to respond readily to pH and salt conditions. Changing these external cues leads to formation or disassembly of aggregates smaller than 100 nm within minutes.
Figures
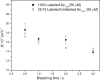
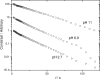
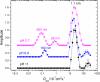
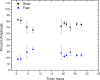
 2.7,
2.7,  6.9, and ■ 11. The vertical scales at pH 6.9 and pH 2.7 have been offset for clarity. Apparent hydrodynamic radii are indicated (does not imply spherical shape for the diffusers.)
6.9, and ■ 11. The vertical scales at pH 6.9 and pH 2.7 have been offset for clarity. Apparent hydrodynamic radii are indicated (does not imply spherical shape for the diffusers.)


Similar articles
-
Amyloid-β oligomerization monitored by single-molecule stepwise photobleaching.Methods. 2021 Sep;193:80-95. doi: 10.1016/j.ymeth.2020.06.007. Epub 2020 Jun 13. Methods. 2021. PMID: 32544592 Free PMC article.
-
Thioflavin T templates amyloid β(1-40) conformation and aggregation pathway.Biophys Chem. 2015 Nov;206:1-11. doi: 10.1016/j.bpc.2015.06.006. Epub 2015 Jun 16. Biophys Chem. 2015. PMID: 26100600
-
Beta-amyloid protein aggregation.Methods Mol Biol. 2007;386:203-25. doi: 10.1007/1-59745-430-3_7. Methods Mol Biol. 2007. PMID: 18604947
-
Elucidating the Structures of Amyloid Oligomers with Macrocyclic β-Hairpin Peptides: Insights into Alzheimer's Disease and Other Amyloid Diseases.Acc Chem Res. 2018 Mar 20;51(3):706-718. doi: 10.1021/acs.accounts.7b00554. Epub 2018 Mar 6. Acc Chem Res. 2018. PMID: 29508987 Free PMC article. Review.
-
Inhibition of the Self-Assembly of Aβ and of Tau by Polyphenols: Mechanistic Studies.Molecules. 2019 Jun 22;24(12):2316. doi: 10.3390/molecules24122316. Molecules. 2019. PMID: 31234523 Free PMC article. Review.
Cited by
-
Biochemistry of amyloid β-protein and amyloid deposits in Alzheimer disease.Cold Spring Harb Perspect Med. 2012 Jun;2(6):a006262. doi: 10.1101/cshperspect.a006262. Cold Spring Harb Perspect Med. 2012. PMID: 22675658 Free PMC article. Review.
-
Characterization of the interaction of β-amyloid with transthyretin monomers and tetramers.Biochemistry. 2010 Sep 28;49(38):8276-89. doi: 10.1021/bi101280t. Biochemistry. 2010. PMID: 20795734 Free PMC article.
-
FRAP in pharmaceutical research: practical guidelines and applications in drug delivery.Pharm Res. 2014 Feb;31(2):255-70. doi: 10.1007/s11095-013-1146-9. Epub 2013 Sep 10. Pharm Res. 2014. PMID: 24019022 Review.
-
Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing.Nat Rev Mol Cell Biol. 2021 Mar;22(3):196-213. doi: 10.1038/s41580-020-00326-6. Epub 2021 Jan 28. Nat Rev Mol Cell Biol. 2021. PMID: 33510441 Review.
-
Aβ40 Aggregation under Changeable Conditions.Int J Mol Sci. 2023 May 7;24(9):8408. doi: 10.3390/ijms24098408. Int J Mol Sci. 2023. PMID: 37176115 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

