Detection of tumor epidermal growth factor receptor pathway dependence by serum mass spectrometry in cancer patients
- PMID: 20086114
- PMCID: PMC2846615
- DOI: 10.1158/1055-9965.EPI-09-0937
Detection of tumor epidermal growth factor receptor pathway dependence by serum mass spectrometry in cancer patients
Abstract
Background: We hypothesized that a serum proteomic profile predictive of survival benefit in non-small cell lung cancer patients treated with epidermal growth factor receptor tyrosine kinase inhibitors (EGFR-TKI) reflects tumor EGFR dependency regardless of site of origin or class of therapeutic agent.
Methods: Pretreatment serum or plasma from 230 patients treated with cetuximab, EGFR-TKIs, or chemotherapy for recurrent/metastatic head and neck squamous cell carcinoma (HNSCC) or colorectal cancer (CRC) were analyzed by mass spectrometry. Each sample was classified into "good" or "poor" groups using VeriStrat, and survival analyses of each cohort were done based on this classification. For the CRC cohort, this classification was correlated with the tumor EGFR ligand levels and KRAS mutation status.
Results: In the EGFR inhibitor-treated cohorts, the classification predicted survival (HNSCC: gefitinib, P = 0.007 and erlotinib/bevacizumab, P = 0.02; CRC: cetuximab, P = 0.0065) whereas the chemotherapy cohort showed no survival difference. For CRC patients, tumor EGFR ligand RNA levels were significantly associated with the proteomic classification, and combined KRAS and proteomic classification provided improved survival classification.
Conclusions: Serum proteomic profiling can detect clinically significant tumor dependence on the EGFR pathway in non-small cell lung cancer, HNSCC, and CRC patients treated with either EGFR-TKIs or cetuximab. This classification is correlated with tumor EGFR ligand levels and provides a clinically practical way to identify patients with diverse cancer types most likely to benefit from EGFR inhibitors. Prospective studies are necessary to confirm these findings.
Figures
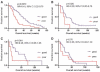
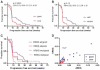
References
-
- Shepherd FA, Rodrigues Pereira J, Ciuleanu T, et al. Erlotinib in previously treated non-small-cell lung cancer. N Engl J Med. 2005;353:123–32. - PubMed
-
- Jonker DJ, O'Callaghan CJ, Karapetis CS, et al. Cetuximab for the treatment of colorectal cancer. N Engl J Med. 2007;357:2040–8. - PubMed
-
- Cohen EE. Role of epidermal growth factor receptor pathway-targeted therapy in patients with recurrent and/or metastatic squamous cell carcinoma of the head and neck. J Clin Oncol. 2006;24:2659–65. - PubMed
-
- Chung KY, Shia J, Kemeny NE, et al. Cetuximab shows activity in colorectal cancer patients with tumors that do not express the epidermal growth factor receptor by immunohistochemistry. J Clin Oncol. 2005;23:1803–10. - PubMed
-
- Van Cutsem E, Peeters M, Siena S, et al. Open-label phase III trial of panitumumab plus best supportive care compared with best supportive care alone in patients with chemotherapy-refractory meta-static colorectal cancer. J Clin Oncol. 2007;25:1658–64. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- U10 CA027525/CA/NCI NIH HHS/United States
- CA66636/CA/NCI NIH HHS/United States
- U10 CA021115/CA/NCI NIH HHS/United States
- CA27525/CA/NCI NIH HHS/United States
- U10 CA017145/CA/NCI NIH HHS/United States
- R01 DE017982/DE/NIDCR NIH HHS/United States
- U10 CA066636/CA/NCI NIH HHS/United States
- CA49957/CA/NCI NIH HHS/United States
- U10 CA016116/CA/NCI NIH HHS/United States
- P30 CA068485/CA/NCI NIH HHS/United States
- U10 CA049957/CA/NCI NIH HHS/United States
- CA16116/CA/NCI NIH HHS/United States
- UL1 TR000005/TR/NCATS NIH HHS/United States
- R01-DE-017982/DE/NIDCR NIH HHS/United States
- U10 CA039229/CA/NCI NIH HHS/United States
- CA39229/CA/NCI NIH HHS/United States
- CA17145/CA/NCI NIH HHS/United States
- U10 CA023318/CA/NCI NIH HHS/United States
- N01-CM-17102/CM/NCI NIH HHS/United States
- CA21115/CA/NCI NIH HHS/United States
- CA068485/CA/NCI NIH HHS/United States
- U24 CA114737/CA/NCI NIH HHS/United States
- CA23318/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

