Deep sequencing of small RNA libraries reveals dynamic regulation of conserved and novel microRNAs and microRNA-stars during silkworm development
- PMID: 20089182
- PMCID: PMC2824724
- DOI: 10.1186/1471-2164-11-52
Deep sequencing of small RNA libraries reveals dynamic regulation of conserved and novel microRNAs and microRNA-stars during silkworm development
Abstract
Background: In eukaryotes, microRNAs (miRNAs) have emerged as critical regulators of gene expression. The Silkworm (Bombyx mori L.) is one of the most suitable lepidopteran insects for studying the molecular aspects of metamorphosis because of its large size, availability of mutants and genome sequence. Besides, this insect also has been amply studied from a physiological and biochemical perspective. Deep sequencing of small RNAs isolated from different stages of silkworm is a powerful tool not only for measuring the changes in miRNA profile but also for discovering novel miRNAs.
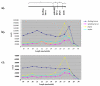
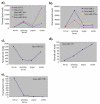
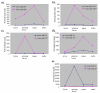
Results: We generated small RNA libraries from feeding larvae, spinning larvae, pupae and adults of B. mori and obtained approximately 2.5 million reads of 18-30 nt. Sequence analysis identified 14 novel and 101 conserved miRNAs. Most novel miRNAs are preferentially expressed in pupae, whereas more than 95% of the conserved miRNAs are dynamically regulated during different developmental stages. Remarkably, the miRNA-star (miR*) of four miRNAs are expressed at much higher levels than their corresponding miRNAs, and their expression profiles are distinct from their corresponding miRNA profiles during different developmental stages. Additionally, we detected two antisense miRNA loci (miR-263-S and miR-263-AS; miR-306-S and miR-306-AS) that are expressed in sense and antisense directions. Interestingly, miR-263 and miR-306 are preferentially and abundantly expressed in pupae and adults, respectively.
Conclusions: We identified 101 homologs of conserved miRNAs, 14 species-specific and two antisense miRNAs in the silkworm. Our results provided deeper insights into changes in conserved and novel miRNA and miRNA* accumulation during development.
Figures




Similar articles
-
Deep sequencing of small RNA libraries reveals dynamic expression patterns of microRNAs in multiple developmental stages of Bactrocera dorsalis.Insect Mol Biol. 2014 Oct;23(5):656-67. doi: 10.1111/imb.12111. Epub 2014 Jun 23. Insect Mol Biol. 2014. PMID: 24957108
-
MicroRNAs of Bombyx mori identified by Solexa sequencing.BMC Genomics. 2010 Mar 3;11:148. doi: 10.1186/1471-2164-11-148. BMC Genomics. 2010. PMID: 20199675 Free PMC article.
-
Insect-Specific microRNA Involved in the Development of the Silkworm Bombyx mori.PLoS One. 2009;4(3):e4677. doi: 10.1371/journal.pone.0004677. Epub 2009 Mar 5. PLoS One. 2009. PMID: 19262741 Free PMC article.
-
Identification of microRNAs from Plutella xylostella larvae associated with parasitization by Diadegma semiclausum.Insect Biochem Mol Biol. 2013 Apr;43(4):309-18. doi: 10.1016/j.ibmb.2013.01.004. Epub 2013 Jan 23. Insect Biochem Mol Biol. 2013. PMID: 23352895 Review.
-
Viral-Encoded microRNAs in Host-Pathogen Interactions in Silkworm.Microrna. 2021;10(1):3-13. doi: 10.2174/2211536610666210121154314. Microrna. 2021. PMID: 33475082 Review.
Cited by
-
Genome-wide analysis of mir-548 gene family reveals evolutionary and functional implications.J Biomed Biotechnol. 2012;2012:679563. doi: 10.1155/2012/679563. Epub 2012 Oct 2. J Biomed Biotechnol. 2012. PMID: 23091353 Free PMC article.
-
Selective activation of miRNAs of the primate-specific chromosome 19 miRNA cluster (C19MC) in cancer and stem cells and possible contribution to regulation of apoptosis.J Biomed Sci. 2017 Mar 7;24(1):20. doi: 10.1186/s12929-017-0326-z. J Biomed Sci. 2017. PMID: 28270145 Free PMC article.
-
IsomiR expression patterns in canonical and Dicer‑independent microRNAs.Mol Med Rep. 2017 Mar;15(3):1071-1078. doi: 10.3892/mmr.2017.6117. Epub 2017 Jan 13. Mol Med Rep. 2017. PMID: 28098889 Free PMC article.
-
Differentially expressed microRNAs in diapausing versus HCl-treated Bombyx embryos.PLoS One. 2017 Jul 11;12(7):e0180085. doi: 10.1371/journal.pone.0180085. eCollection 2017. PLoS One. 2017. PMID: 28700597 Free PMC article.
-
The Tomato Spotted Wilt Virus Genome Is Processed Differentially in its Plant Host Arachis hypogaea and its Thrips Vector Frankliniella fusca.Front Plant Sci. 2016 Sep 7;7:1349. doi: 10.3389/fpls.2016.01349. eCollection 2016. Front Plant Sci. 2016. PMID: 27656190 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

