Punique virus, a novel phlebovirus, related to sandfly fever Naples virus, isolated from sandflies collected in Tunisia
- PMID: 20089800
- PMCID: PMC3496376
- DOI: 10.1099/vir.0.019240-0
Punique virus, a novel phlebovirus, related to sandfly fever Naples virus, isolated from sandflies collected in Tunisia
Abstract
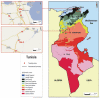
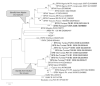
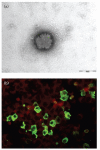
Sandflies are widely distributed around the Mediterranean Basin. Therefore, human populations in this area are potentially exposed to sandfly-transmitted diseases, including those caused by phleboviruses. Whilst there are substantial data in countries located in the northern part of the Mediterranean basin, few data are available for North Africa. In this study, a total of 1489 sandflies were collected in 2008 in Tunisia from two sites, bioclimatically distinct, located 235 km apart, and identified morphologically. Sandfly species comprised Phlebotomus perniciosus (52.2%), Phlebotomus longicuspis (30.1%), Phlebotomus papatasi (12.0%), Phlebotomus perfiliewi (4.6%), Phlebotomus langeroni (0.4%) and Sergentomyia minuta (0.5%). PCR screening, using generic primers for the genus Phlebovirus, resulted in the detection of ten positive pools. Sequence analysis revealed that two pools contained viral RNA corresponding to a novel virus closely related to sandfly fever Naples virus. Virus isolation in Vero cells was achieved from one pool. Genetic and phylogenetic characterization based on sequences in the three genomic segments showed that it was a novel virus distinct from other recognized members of the species. This novel virus was provisionally named Punique virus. Viral sequences in the polymerase gene corresponding to another phlebovirus closely related to but distinct from sandfly fever Sicilian virus were obtained from the eight remaining positive pools.
Figures

References
-
- Chastel C, Bach-Hamba D, Karabatsos N, Bouattour A, Le Lay G, Le Goff F, Vermeil C. Tunis virus: a new Phlebovirus from Argas reflexus hermanni ticks in Tunisia. Acta Virol. 1994;38:285–289. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

