Mycobacteriophage Ms6 LysB specifically targets the outer membrane of Mycobacterium smegmatis
- PMID: 20093291
- PMCID: PMC2889447
- DOI: 10.1099/mic.0.032821-0
Mycobacteriophage Ms6 LysB specifically targets the outer membrane of Mycobacterium smegmatis
Abstract
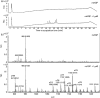
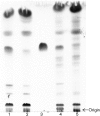
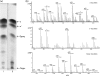
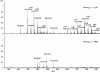
LysB, a mycobacteriophage Ms6-encoded protein, was previously identified as a lipolytic enzyme able to hydrolyse the ester bond in lipase and esterase substrates. In the present work, we show that LysB can hydrolyse lipids containing mycolic acids from the outer membrane of the mycobacterial cell wall. LysB was shown to hydrolyse the mycolic acids from the mycolyl-arabinogalactan-peptidoglycan complex where the mycolates of the inner leaflet of the outer membrane are covalently attached to an arabinosyl head group. In addition, treatment of the extractable lipids from Mycobacterium smegmatis, Mycobacterium bovis BCG and Mycobacterium tuberculosis H37Ra with LysB showed that trehalose 6,6'-dimycolate (TDM), a trehalose diester of two mycolic acid molecules, was hydrolysed by the enzyme. We have also determined the structures of the mycolic acid molecules that form the M. smegmatis TDM. The identification of a phage-encoded enzyme that targets the outer membrane of the mycobacterial cell wall enhances our understanding of the mechanism of mycobacteriophage lysis.
Figures

Similar articles
-
Mycobacteriophage Lysin B is a novel mycolylarabinogalactan esterase.Mol Microbiol. 2009 Aug;73(3):367-81. doi: 10.1111/j.1365-2958.2009.06775.x. Epub 2009 Jun 22. Mol Microbiol. 2009. PMID: 19555454 Free PMC article.
-
The Ms6 Mycolyl-Arabinogalactan Esterase LysB is Essential for an Efficient Mycobacteriophage-Induced Lysis.Viruses. 2017 Nov 17;9(11):343. doi: 10.3390/v9110343. Viruses. 2017. PMID: 29149017 Free PMC article.
-
The Mycobacteriophage Ms6 LysB N-Terminus Displays Peptidoglycan Binding Affinity.Viruses. 2021 Jul 15;13(7):1377. doi: 10.3390/v13071377. Viruses. 2021. PMID: 34372584 Free PMC article.
-
Mycobacteriophage Lysis Enzymes: Targeting the Mycobacterial Cell Envelope.Viruses. 2018 Aug 14;10(8):428. doi: 10.3390/v10080428. Viruses. 2018. PMID: 30110929 Free PMC article. Review.
-
Cell wall synthesizing complexes in Mycobacteriales.Curr Opin Microbiol. 2024 Jun;79:102478. doi: 10.1016/j.mib.2024.102478. Epub 2024 Apr 22. Curr Opin Microbiol. 2024. PMID: 38653035 Review.
Cited by
-
Genomic and functional analyses of Rhodococcus equi phages ReqiPepy6, ReqiPoco6, ReqiPine5, and ReqiDocB7.Appl Environ Microbiol. 2011 Jan;77(2):669-83. doi: 10.1128/AEM.01952-10. Epub 2010 Nov 19. Appl Environ Microbiol. 2011. PMID: 21097585 Free PMC article.
-
Functional analysis of the holin-like proteins of mycobacteriophage Ms6.J Bacteriol. 2011 Jun;193(11):2793-803. doi: 10.1128/JB.01519-10. Epub 2011 Mar 25. J Bacteriol. 2011. PMID: 21441511 Free PMC article.
-
Mycobacteriophages as Potential Therapeutic Agents against Drug-Resistant Tuberculosis.Int J Mol Sci. 2021 Jan 13;22(2):735. doi: 10.3390/ijms22020735. Int J Mol Sci. 2021. PMID: 33450990 Free PMC article. Review.
-
Genome Sequence of Gordonia Phage Emalyn.Genome Announc. 2016 Aug 11;4(4):e00597-16. doi: 10.1128/genomeA.00597-16. Genome Announc. 2016. PMID: 27516499 Free PMC article.
-
Genomic characterization of the antiviral arsenal of Actinobacteria.Microbiology (Reading). 2023 Aug;169(8):001374. doi: 10.1099/mic.0.001374. Microbiology (Reading). 2023. PMID: 37531269 Free PMC article.
References
-
- Barry, C. E., III, Lee, R. E., Mdluli, K., Sampson, A. E., Schroeder, B. G., Slayden, R. A. & Yuan, Y. (1998). Mycolic acids: structure, biosynthesis and physiological functions. Prog Lipid Res 37, 143–179. - PubMed
-
- Bhamidi, S., Scherman, M. S., Rithner, C. D., Prenni, J. E., Chatterjee, D., Khoo, K.-H. & McNeil, M. (2008). The identification and location of succinyl residues and the characterization of the interior arabinan region allow for a model of the complete primary structure of Mycobacterium tuberculosis mycolyl arabinogalactan. J Biol Chem 283, 12992–13000. - PMC - PubMed
-
- Crick, D. C. & Brennan, P. J. (2008). Biosynthesis of the arabinogalactan–peptidoglycan complex of Mycobacterium tuberculosis. In The Mycobacterial Cell Envelope, pp. 25–40. Edited by M. Daffé & J.-M. Reyrat. Washington, DC: American Society for Microbiology.
-
- Daffé, M. (2008). The global architecture of the mycobacterial cell envelope. In The Mycobacterial Cell Envelope, pp. 3–11. Edited by M. Daffé & J.-M. Reyrat. Washington, DC: American Society for Microbiology.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

