Structural insights into de novo actin polymerization
- PMID: 20096561
- PMCID: PMC2854303
- DOI: 10.1016/j.sbi.2009.12.012
Structural insights into de novo actin polymerization
Abstract
Many cellular functions depend on rapid and localized actin polymerization/depolymerization. Yet, the de novo polymerization of actin in cells is kinetically unfavorable because of the instability of polymerization intermediates (small actin oligomers) and the actions of actin monomer binding proteins. Cells use filament nucleation and elongation factors to initiate and sustain polymerization. Structural biology is beginning to shed light on the diverse mechanisms by which these unrelated proteins initiate polymerization, undergo regulation, and mediate the transition of monomeric actin onto actin filaments. A prominent role is played by the W domain, which in some of these proteins occurs in tandem repeats that recruit multiple actin subunits. Pro-rich regions are also abundant and mediate the binding of profilin-actin complexes, which are the main source of polymerization competent actin in cells. Filament nucleation and elongation factors frequently interact with Rho-family GTPases, which relay signals from membrane receptors to regulate actin cytoskeleton remodeling.
Figures

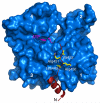



References
-
- Holmes KC. Structural biology: actin in a twist. Nature. 2009;457:389–390. - PubMed
-
- Goode BL, Eck MJ. Mechanism and function of formins in the control of actin assembly. Annu Rev Biochem. 2007;76:593–627. - PubMed
-
- Higgs HN. Formin proteins: a domain-based approach. Trends Biochem Sci. 2005;30:342–353. - PubMed
-
- Castrillon DH, Wasserman SA. Diaphanous is required for cytokinesis in Drosophila and shares domains of similarity with the products of the limb deformity gene. Development. 1994;120:3367–3377. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

