Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency
- PMID: 20098412
- PMCID: PMC2965733
- DOI: 10.1038/nature08829
Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency
Abstract
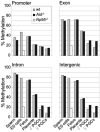
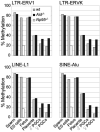
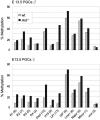
Epigenetic reprogramming including demethylation of DNA occurs in mammalian primordial germ cells (PGCs) and in early embryos, and is important for the erasure of imprints and epimutations, and the return to pluripotency. The extent of this reprogramming and its molecular mechanisms are poorly understood. We previously showed that the cytidine deaminases AID and APOBEC1 can deaminate 5-methylcytosine in vitro and in Escherichia coli, and in the mouse are expressed in tissues in which demethylation occurs. Here we profiled DNA methylation throughout the genome by unbiased bisulphite next generation sequencing in wild-type and AID-deficient mouse PGCs at embryonic day (E)13.5. Wild-type PGCs revealed marked genome-wide erasure of methylation to a level below that of methylation deficient (Np95(-/-), also called Uhrf1(-/-)) embryonic stem cells, with female PGCs being less methylated than male ones. By contrast, AID-deficient PGCs were up to three times more methylated than wild-type ones; this substantial difference occurred throughout the genome, with introns, intergenic regions and transposons being relatively more methylated than exons. Relative hypermethylation in AID-deficient PGCs was confirmed by analysis of individual loci in the genome. Our results reveal that erasure of DNA methylation in the germ line is a global process, hence limiting the potential for transgenerational epigenetic inheritance. AID deficiency interferes with genome-wide erasure of DNA methylation patterns, indicating that AID has a critical function in epigenetic reprogramming and potentially in restricting the inheritance of epimutations in mammals.
Figures

References
-
- Reik W, Dean W, Walter J. Epigenetic reprogramming in mammalian development. Science. 2001;293:1089–1093. - PubMed
-
- Sasaki H, Matsui Y. Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nature Rev Genet. 2008;9:129–140. - PubMed
-
- Oswald J, et al. Active demethylation of the paternal genome in the mouse zygote. Curr Biol. 2000;10:475–478. - PubMed
-
- Mayer W, Niveleau A, Walter J, Fundele R, Haaf T. Demethylation of the zygotic paternal genome. Nature. 2000;403:501–501. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
- R37 GM060398/GM/NIGMS NIH HHS/United States
- G0700098/MRC_/Medical Research Council/United Kingdom
- HHMI/Howard Hughes Medical Institute/United States
- BBS/E/B/0000M220/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BBS/E/B/0000M982/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

