Structural insights into the prereaction state of pyruvate decarboxylase from Zymomonas mobilis
- PMID: 20099870
- PMCID: PMC2855724
- DOI: 10.1021/bi901864j
Structural insights into the prereaction state of pyruvate decarboxylase from Zymomonas mobilis
Abstract
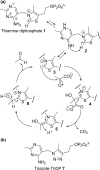
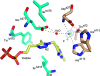
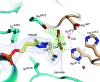
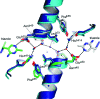
Pyruvate decarboxylase (PDC) uses thiamine diphosphate as an essential cofactor to catalyze the formation of acetaldehyde on the pathway of ethanol synthesis. Here we report the crystallographic image of a prereaction intermediate of a bacterial pyruvate decarboxylase prepared by cocrystallizing the enzyme with pyruvate and a stable analogue of the cofactor's activated ylid form. A second crystal structure of PDC in complex with fluoride shows that the ion organizes a water molecule that occludes the pyruvate binding site, accounting for the inhibitory effect of the halide. Also reported is a structure of the cofactor-free apo form, which when compared to the structure of the holo form indicates how thiamine diphosphate organizes the active site pocket of pyruvate decarboxylase to support catalysis. Guided by the structural and enzymatic data, we propose roles for several key residues in the catalytic mechanism.
Figures
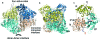
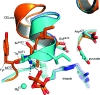

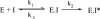
Similar articles
-
Structure and properties of pyruvate decarboxylase and site-directed mutagenesis of the Zymomonas mobilis enzyme.Biochim Biophys Acta. 1998 Jun 29;1385(2):323-38. doi: 10.1016/s0167-4838(98)00077-6. Biochim Biophys Acta. 1998. PMID: 9655927 Review.
-
High resolution crystal structure of pyruvate decarboxylase from Zymomonas mobilis. Implications for substrate activation in pyruvate decarboxylases.J Biol Chem. 1998 Aug 7;273(32):20196-204. doi: 10.1074/jbc.273.32.20196. J Biol Chem. 1998. PMID: 9685367
-
Mutagenesis at asp27 of pyruvate decarboxylase from Zymomonas mobilis. Effect on its ability to form acetoin and acetolactate.Eur J Biochem. 2000 Nov;267(21):6493-500. doi: 10.1046/j.1432-1327.2000.01744.x. Eur J Biochem. 2000. PMID: 11029594
-
The role of His113 and His114 in pyruvate decarboxylase from Zymomonas mobilis.Eur J Biochem. 1997 Aug 15;248(1):63-71. doi: 10.1111/j.1432-1033.1997.t01-1-00063.x. Eur J Biochem. 1997. PMID: 9310361
-
Subunit structure, function and organisation of pyruvate decarboxylases from various organisms.Biochim Biophys Acta. 1998 Jun 29;1385(2):271-86. doi: 10.1016/s0167-4838(98)00074-0. Biochim Biophys Acta. 1998. PMID: 9655918 Review.
Cited by
-
Fluoride-mediated capture of a noncovalent bound state of a reversible covalent enzyme inhibitor: X-ray crystallographic analysis of an exceptionally potent α-ketoheterocycle inhibitor of fatty acid amide hydrolase.J Am Chem Soc. 2011 Mar 23;133(11):4092-100. doi: 10.1021/ja110877y. Epub 2011 Feb 28. J Am Chem Soc. 2011. PMID: 21355555 Free PMC article.
-
An iterative computational design approach to increase the thermal endurance of a mesophilic enzyme.Biotechnol Biofuels. 2018 Jul 9;11:189. doi: 10.1186/s13068-018-1178-9. eCollection 2018. Biotechnol Biofuels. 2018. PMID: 30002729 Free PMC article.
-
Experimental observation of thiamin diphosphate-bound intermediates on enzymes and mechanistic information derived from these observations.Bioorg Chem. 2005 Jun;33(3):190-215. doi: 10.1016/j.bioorg.2005.02.001. Epub 2005 Apr 1. Bioorg Chem. 2005. PMID: 15888311 Free PMC article. Review.
-
Crystal structure of an inferred ancestral bacterial pyruvate decarboxylase.Acta Crystallogr F Struct Biol Commun. 2018 Mar 1;74(Pt 3):179-186. doi: 10.1107/S2053230X18002819. Epub 2018 Feb 26. Acta Crystallogr F Struct Biol Commun. 2018. PMID: 29497023 Free PMC article.
-
Inhibition of Thiamine Diphosphate-Dependent Enzymes by Triazole-Based Thiamine Analogues.ACS Med Chem Lett. 2023 Apr 11;14(5):621-628. doi: 10.1021/acsmedchemlett.3c00047. eCollection 2023 May 11. ACS Med Chem Lett. 2023. PMID: 37197459 Free PMC article.
References
-
- Jordan F. (2003) Current mechanistic understanding of thiamin diphosphate-dependent enzymatic reactions. Nat. Prod. Rep. 20, 184–201. - PubMed
-
- Schellenberger A. (1998) Sixty years of thiamin diphosphate biochemistry. Biochim. Biophys. Acta 1385, 177–186. - PubMed
-
- Kern D.; Kern G.; Neef H.; Tittmann K.; Killenberg-Jabs M.; Wikner C.; Schneider G.; Hübner G. (1997) How Thiamin Diphosphate Is Activated in Enzymes. Science 275, 67–70. - PubMed
-
- Nemeria N.; Korotchkina L.; McLeish M. J.; Kenyon G. L.; Patel M. S.; Jordan F. (2007) Elucidation of the Chemistry of Enzyme-Bound Thiamin Diphosphate Prior to Substrate Binding: Defining Internal Equilibria among Tautomeric and Ionization States. Biochemistry 46, 10739–10744. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

