Kinetics of PIP2 metabolism and KCNQ2/3 channel regulation studied with a voltage-sensitive phosphatase in living cells
- PMID: 20100891
- PMCID: PMC2812502
- DOI: 10.1085/jgp.200910345
Kinetics of PIP2 metabolism and KCNQ2/3 channel regulation studied with a voltage-sensitive phosphatase in living cells
Abstract
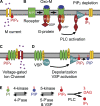
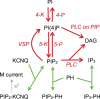
The signaling phosphoinositide phosphatidylinositol 4,5-bisphosphate (PIP(2)) is synthesized in two steps from phosphatidylinositol by lipid kinases. It then interacts with KCNQ channels and with pleckstrin homology (PH) domains among many other physiological protein targets. We measured and developed a quantitative description of these metabolic and protein interaction steps by perturbing the PIP(2) pool with a voltage-sensitive phosphatase (VSP). VSP can remove the 5-phosphate of PIP(2) with a time constant of tau <300 ms and fully inhibits KCNQ currents in a similar time. PIP(2) was then resynthesized from phosphatidylinositol 4-phosphate (PIP) quickly, tau = 11 s. In contrast, resynthesis of PIP(2) after activation of phospholipase C by muscarinic receptors took approximately 130 s. These kinetic experiments showed that (1) PIP(2) activation of KCNQ channels obeys a cooperative square law, (2) the PIP(2) residence time on channels is <10 ms and the exchange time on PH domains is similarly fast, and (3) the step synthesizing PIP(2) by PIP 5-kinase is fast and limited primarily by a step(s) that replenishes the pool of plasma membrane PI(4)P. We extend the kinetic model for signaling from M(1) muscarinic receptors, presented in our companion paper in this issue (Falkenburger et al. 2010. J. Gen. Physiol. doi:10.1085/jgp.200910344), with this new information on PIP(2) synthesis and KCNQ interaction.
Figures



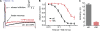
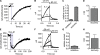


References
-
- Balla A., Tuymetova G., Tsiomenko A., Várnai P., Balla T. 2005. A plasma membrane pool of phosphatidylinositol 4-phosphate is generated by phosphatidylinositol 4-kinase type-III alpha: studies with the PH domains of the oxysterol binding protein and FAPP1. Mol. Biol. Cell. 16:1282–1295 10.1091/mbc.E04-07-0578 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

