Stringent requirement for HRD1, SEL1L, and OS-9/XTP3-B for disposal of ERAD-LS substrates
- PMID: 20100910
- PMCID: PMC2812524
- DOI: 10.1083/jcb.200910042
Stringent requirement for HRD1, SEL1L, and OS-9/XTP3-B for disposal of ERAD-LS substrates
Abstract
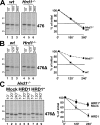
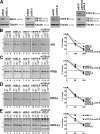
Sophisticated quality control mechanisms prolong retention of protein-folding intermediates in the endoplasmic reticulum (ER) until maturation while sorting out terminally misfolded polypeptides for ER-associated degradation (ERAD). The presence of structural lesions in the luminal, transmembrane, or cytosolic domains determines the classification of misfolded polypeptides as ERAD-L, -M, or -C substrates and results in selection of distinct degradation pathways. In this study, we show that disposal of soluble (nontransmembrane) polypeptides with luminal lesions (ERAD-L(S) substrates) is strictly dependent on the E3 ubiquitin ligase HRD1, the associated cargo receptor SEL1L, and two interchangeable ERAD lectins, OS-9 and XTP3-B. These ERAD factors become dispensable for degradation of the same polypeptides when membrane tethered (ERAD-L(M) substrates). Our data reveal that, in contrast to budding yeast, tethering of mammalian ERAD-L substrates to the membrane changes selection of the degradation pathway.
Figures
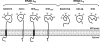


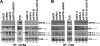

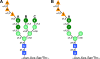
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

